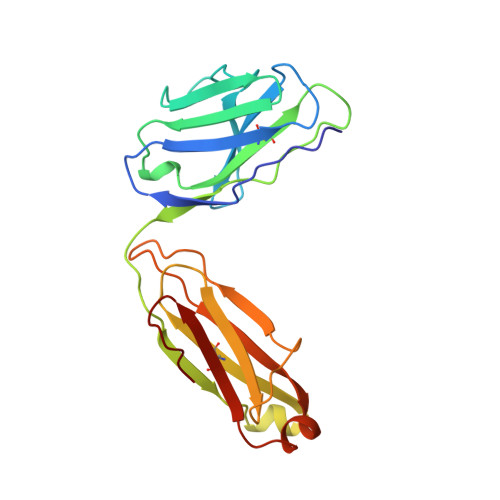
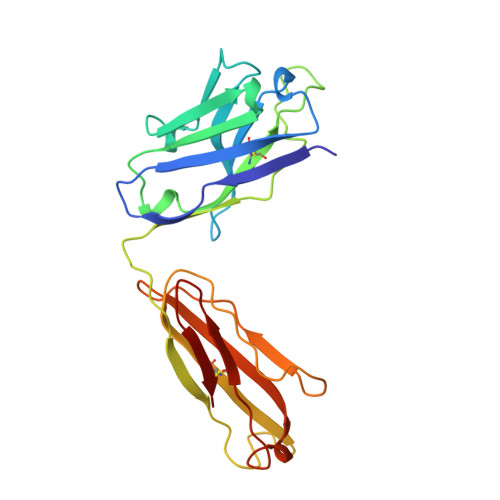
The role of homophilic binding in anti-tumor antibody R24 recognition of molecular surfaces. Demonstration of an intermolecular beta-sheet interaction between vh domains.
Kaminski, M.J., MacKenzie, C.R., Mooibroek, M.J., Dahms, T.E., Hirama, T., Houghton, A.N., Chapman, P.B., Evans, S.V.(1999) J Biological Chem 274: 5597-5604
- PubMed: 10026176 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.274.9.5597
- Primary Citation Related Structures:
1BZ7, 1R24 - PubMed Abstract:
The murine antibody R24 and mouse-human Fv-IgG1(kappa) chimeric antibody chR24 are specific for the cell-surface tumor antigen disialoganglioside GD3. X-ray diffraction and surface plasmon resonance experiments have been employed to study the mechanism of "homophilic binding," in which molecules of R24 recognize and bind to other molecules of R24 though their heavy chain variable domains. R24 exhibits strong binding to liposomes containing disialoganglioside GD3; however, the kinetics are unusual in that saturation of binding is not observed. The binding of chR24 to GD3-bearing liposomes is significantly weaker, suggesting that cooperative interactions involving antibody constant regions contribute to R24 binding of membrane-bound GD3. The crystal structures of the Fabs from R24 and chR24 reveal the mechanism for homophilic binding and confirm that the homophilic and antigen-binding idiotopes are distinct. The homophilic binding idiotope is formed largely by an anti-parallel beta-sheet dimerization between the H2 complementarity determining region (CDR) loops of two Fabs, while the antigen-binding idiotope is a pocket formed by the three CDR loops on the heavy chain. The formation of homophilic dimers requires the presence of a canonical conformation for the H2 CDR in conjunction with participation of side chains. The relative positions of the homophilic and antigen-binding sites allows for a lattice of GD3-specific antibodies to be constructed, which is stabilized by the presence of the cell membrane. This model provides for the selective recognition by R24 of cells that overexpress GD3 on the cell surface.
- Department of Biochemistry, University of Ottawa, Ottawa, Ontario K1H 8M5, Canada.
Organizational Affiliation: