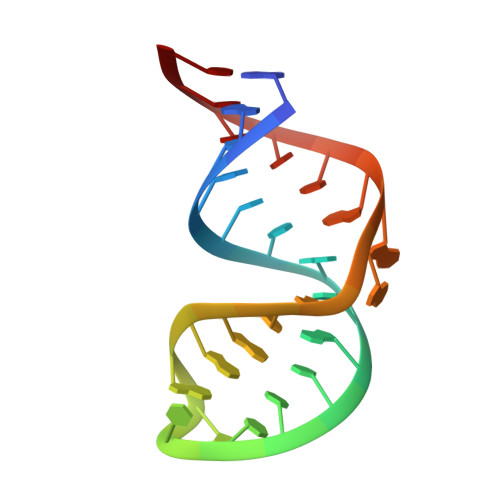
Structural origins of gentamicin antibiotic action.
Yoshizawa, S., Fourmy, D., Puglisi, J.D.(1998) EMBO J 17: 6437-6448
- PubMed: 9822590 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/17.22.6437
- Primary Citation Related Structures:
1BYJ - PubMed Abstract:
Aminoglycoside antibiotics that bind to the ribosomal A site cause misreading of the genetic code and inhibit translocation. The clinically important aminoglycoside, gentamicin C, is a mixture of three components. Binding of each gentamicin component to the ribosome and to a model RNA oligonucleotide was studied biochemically and the structure of the RNA complexed to gentamicin C1a was solved using magnetic resonance nuclear spectroscopy. Gentamicin C1a binds in the major groove of the RNA. Rings I and II of gentamicin direct specific RNA-drug interactions. Ring III of gentamicin, which distinguishes this subclass of aminoglycosides, also directs specific RNA interactions with conserved base pairs. The structure leads to a general model for specific ribosome recognition by aminoglycoside antibiotics and a possible mechanism for translational inhibition and miscoding. This study provides a structural rationale for chemical synthesis of novel aminoglycosides.
- Department of Structural Biology, Stanford University School of Medicine, Stanford, CA 94305-5400, USA.
Organizational Affiliation: