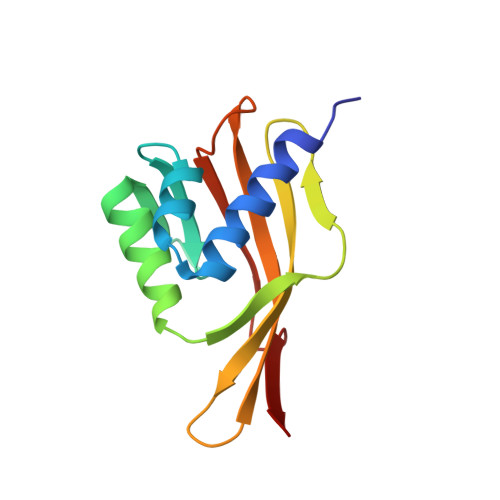
Solution structure of Delta 5-3-ketosteroid isomerase complexed with the steroid 19-nortestosterone hemisuccinate.
Massiah, M.A., Abeygunawardana, C., Gittis, A.G., Mildvan, A.S.(1998) Biochemistry 37: 14701-14712
- PubMed: 9778345 Search on PubMed
- DOI: https://doi.org/10.1021/bi981447b
- Primary Citation Related Structures:
1BUQ - PubMed Abstract:
The solution structure of the ketosteroid isomerase homodimer complexed with the product analogue 19-nortestosterone hemisuccinate (19-NTHS) was solved by heteronuclear multidimensional NMR methods using 1647 distance restraints, 77 dihedral angle (phi) restraints, and 67 hydrogen bond restraints per monomer. The refined secondary structure of each subunit consists of three alpha-helices, eight beta-strands, four turns, and two beta-bulges. The beta-strands form a mixed beta-sheet. One of the five proline residues, Pro-39, is cis and begins a nonclassical turn. A self-consistent ensemble of 15 tertiary/quaternary structures of the enzyme dimer-steroid complex, with no distance violations greater than 0.35 A, was generated by simulated annealing and energy minimization with the program X-PLOR. The mean pairwise RMSD of the secondary structural elements was 0.63 A for the average subunit and 1.25 A for the dimer. Within each subunit, the three alpha-helices are packed onto the concave surface of the beta-sheet with a groove between them into which the steroid binds at a site defined by 14 intermolecular distances. In the productive complex, Tyr-14, from alpha-helix 1, approaches both Asp-99 and the 3-keto group of 19-NTHS while, from beta-strand 1, the carboxylate of Asp-38 approaches the beta-face of the steroid near C4 and C6, between which it transfers a proton during catalysis. Thus the solution structure of the isomerase-steroid complex can accommodate the catalytic diad mechanism in which Asp-99 donates a hydrogen bond to Tyr-14 which in turn is hydrogen bonded to the 3-oxygen of the steroid. While direct hydrogen bonding of Asp-99 to the steroid oxygen is less likely, it cannot be excluded. All other interactions of the steroid with the enzyme are hydrophobic. The dimer interface, which is between the convex surfaces of the beta-sheets, is defined by 28 intersubunit NOEs between hydrophobic residues in the 13C-filtered NOESY-HSQC spectrum of a 13C/12C-heterolabeled dimer. Both hydrophobic and polar interactions occur at the dimer interface which contains no space that would permit additional steroid binding. Comparison of the complexed enzyme with the solution structure of the free enzyme [Wu et al. (1997) Science 276, 415-418] reveals that the three helices change position in the steroid complex, becoming more closely packed onto the concave surface of the beta-sheet, thus bringing Tyr-14 closer to Asp-99 and the substrate. Comparison of the enzyme-steroid complex in solution with the free enzyme in the crystalline state reveals similar differences between the positions of the helices.
- Department of Biological Chemistry, The Johns Hopkins University School of Medicine, Baltimore, Maryland 21205, USA.
Organizational Affiliation: