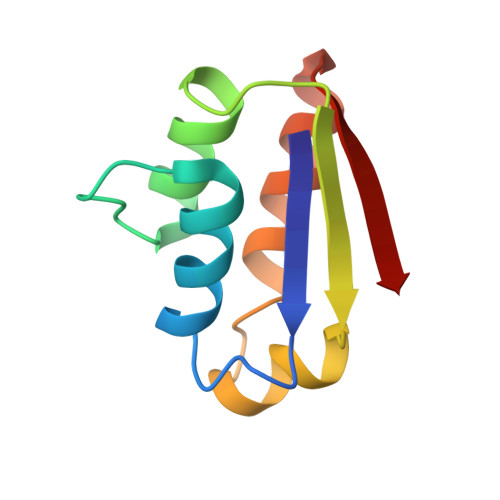
Three-dimensional solution structure and 13C assignments of barstar using nuclear magnetic resonance spectroscopy.
Lubienski, M.J., Bycroft, M., Freund, S.M., Fersht, A.R.(1994) Biochemistry 33: 8866-8877
- PubMed: 8043574 Search on PubMed
- Primary Citation Related Structures:
1BTA, 1BTB - PubMed Abstract:
We present the high-resolution solution structure and 13C assignments of wild-type barstar, an 89 amino acid residue polypeptide inhibitor of barnase, derived from heteronuclear NMR techniques. These were obtained from measurements on unlabeled, uniformly 15N- and 13C/15N-labeled, and 10% 13C-labeled barstar samples that have both cysteines (at positions 40 and 82) fully reduced. In total, 30 structures were calculated by hybrid distance geometry-dynamical simulated annealing calculations. The atomic rms distribution about the mean coordinate positions is 0.42 A for all backbone atoms and 0.90 A for all atoms. The structure is composed of three parallel alpha-helices packed against a three-stranded parallel beta-sheet. A more poorly defined helix links the second beta-strand and the third major alpha-helix. The loop involved in binding barnase is extremely well defined and held rigidly by interactions from the main body of the protein to both ends and the middle of the loop. This structure will be used to aid protein engineering studies currently taking place on the free and bound states of barstar and barnase.
- Cambridge Centre for Protein Engineering, Department of Chemistry, University of Cambridge, U.K.
Organizational Affiliation: