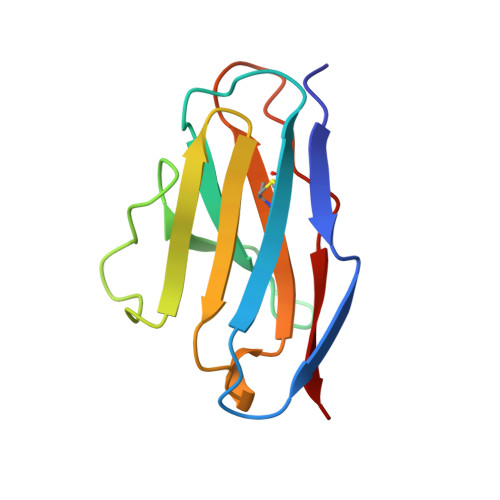
Tertiary structure of an amyloid immunoglobulin light chain protein: a proposed model for amyloid fibril formation.
Schormann, N., Murrell, J.R., Liepnieks, J.J., Benson, M.D.(1995) Proc Natl Acad Sci U S A 92: 9490-9494
- PubMed: 7568160 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.92.21.9490
- Primary Citation Related Structures:
1BRE - PubMed Abstract:
An immunoglobulin light chain protein was isolated from the urine of an individual (BRE) with systemic amyloidosis. Complete amino acid sequence of the variable region of the light chain (VL) protein established it as a kappa I, which when compared with other kappa I amyloid associated proteins had unique residues, including Ile-34, Leu-40, and Tyr-71. To study the tertiary structure, BRE VL was expressed in Escherichia coli by using a PCR product amplified from the patient BRE's bone marrow DNA. The PCR product was ligated into pCZ11, a thermal-inducible replication vector. Recombinant BRE VL was isolated, purified to homogeneity, and crystallized by using ammonium sulfate as the precipitant. Two crystal forms were obtained. In crystal form I the BRE VL kappa domain crystallizes as a dimer with unit cell constants isomorphous to previously published kappa protein structures. Comparison with a nonamyloid VL kappa domain from patient REI, identified significant differences in position of residues in the hypervariable segments plus variations in framework region (FR) segments 40-46 (FR2) and 66-67 (FR3). In addition, positional differences can be seen along the two types of local diads, corresponding to the monomer-monomer and dimer-dimer interfaces. From the packing diagram, a model for the amyloid light chain (AL) fibril is proposed based on a pseudohexagonal spiral structure with a rise of approximately the width of two dimers per 360 degree turn. This spiral structure could be consistent with the dimensions of amyloid fibrils as determined by electron microscopy.
- Department of Medicine, Indiana University School of Medicine, Indianapolis 46202, USA.
Organizational Affiliation: