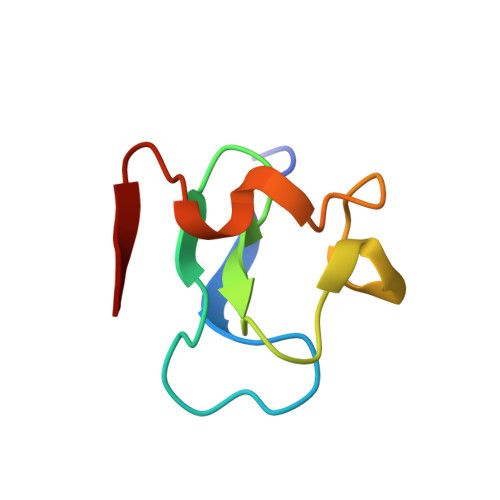
The solution structure of the RING finger domain from the acute promyelocytic leukaemia proto-oncoprotein PML.
Borden, K.L., Boddy, M.N., Lally, J., O'Reilly, N.J., Martin, S., Howe, K., Solomon, E., Freemont, P.S.(1995) EMBO J 14: 1532-1541
- PubMed: 7729428 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1995.tb07139.x
- Primary Citation Related Structures:
1BOR - PubMed Abstract:
Acute promyelocytic leukaemia (APL) has been ascribed to a chromosomal translocation event which results in a fusion protein comprising the PML protein and the retinoic acid receptor alpha. PML is normally a component of a nuclear multiprotein complex (termed ND10, Kr bodies, nuclear bodies, PML oncogenic domains or PODs) which is disrupted in the APL disease state. PML contains a number of characterized motifs including a Zn2+ binding domain called the RING or C3HC4 finger. Here we describe the solution structure of the PML RING finger as solved by 1H NMR methods at physiological pH with r.m.s. deviations for backbone atoms of 0.88 and 1.39 A for all atoms. Additional biophysical studies including CD and optical spectroscopy, show that the PML RING finger requires Zn2+ for autonomous folding and that cysteines are used in metal ligation. A comparison of the structure with the previously solved equine herpes virus IE110 RING finger, shows significant differences suggesting that the RING motif is structurally diverse. The role of the RING domain in PML nuclear body formation was tested in vivo, by using site-directed mutagenesis and immunofluorescence on transiently transfected NIH 3T3 cells. Independently mutating two pairs of cysteines in each of the Zn2+ binding sites prevents PML nuclear body formation, suggesting that a fully folded RING domain is necessary for this process. These results suggest that the PML RING domain is probably involved in protein-protein interactions, a feature which may be common to other RING finger domains.
- Laboratory of Molecular Structure, Biochemistry, National Institute for Medical Research, Mill Hill, London, UK.
Organizational Affiliation: