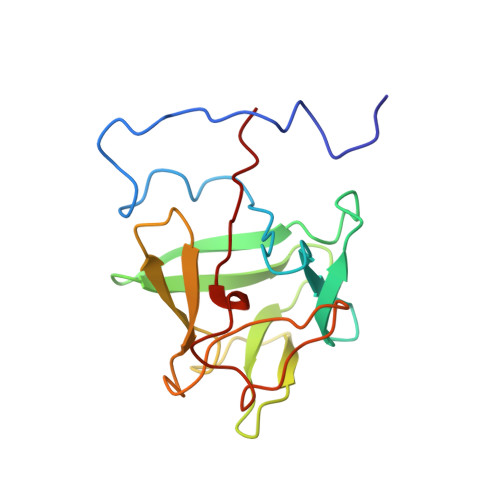
High-resolution solution structure of basic fibroblast growth factor determined by multidimensional heteronuclear magnetic resonance spectroscopy.
Moy, F.J., Seddon, A.P., Bohlen, P., Powers, R.(1996) Biochemistry 35: 13552-13561
- PubMed: 8885834 Search on PubMed
- DOI: https://doi.org/10.1021/bi961260p
- Primary Citation Related Structures:
1BLA, 1BLD - PubMed Abstract:
The high-resolution solution structure of recombinant human basic fibroblast growth factor (FGF-2), a protein of 17.2 kDa that exhibits a variety of functions related to cell growth and differentiation, has been determined using three-dimensional heteronuclear NMR spectroscopy. A total of 30 structures were calculated by means of hybrid distance geometry--simulated annealing using a total of 2865 experimental NMR restraints, consisting of 2486 approximate inteproton distance restraints, 50 distance restraints for 25 backbone hydrogen bonds, and 329 torsion angle restraints. The atomic rms distribution about the mean coordinate positions for the 30 structures for residues 29-152 is 0.43 +/- 0.03 A for the backbone atoms, 0.83 +/- 0.05 A for all atoms, and 0.51 +/- 0.04 A for all atoms excluding disordered side chains. The overall structure of FGF-2 consists of 11 extended antiparallel beta-strands arranged in three groups of three or four strands connected by tight turns and loop regions creating a pseudo-3-fold symmetry. Two strands from each group come together to form a beta-sheet barrel of six antiparallel beta-strands. A helix-like structure was observed for residues 131-136, which is part of the heparin binding site (residues 128-138). The discovery of the helix-like region in the primary heparin binding site instead of the beta-strand conformation described in the X-ray structures may have important implications in understanding the nature of heparin--FGF-2 interactions. A total of seven tightly bound water molecules were found in the FGF-2 structure, two of which are located in the heparin binding site. The first 28 N-terminal residues appear to be disordered, which is consistent with previous X-ray structures. A best fit superposition of the NMR structure of FGF-2 with the 1.9 A resolution X-ray structure by Zhu et al. (1991) yields a backbone atomic rms difference of 0.94 A, indicative of a close similarity between the NMR and X-ray structures.
- Department of Structural Biology, Wyeth-Ayerst Research, Pearl River, New York 10965, USA.
Organizational Affiliation: