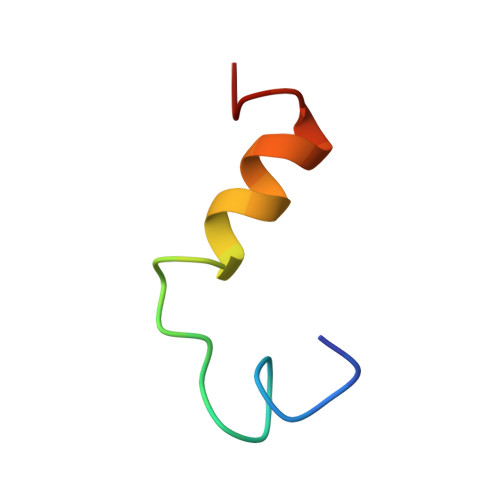
Solution structures in aqueous SDS micelles of two amyloid beta peptides of A beta(1-28) mutated at the alpha-secretase cleavage site (K16E, K16F)
Poulsen, S.-A., Watson, A.A., Craik, D.J.(2000) J Struct Biol 130: 142-152
- PubMed: 10940222 Search on PubMed
- DOI: https://doi.org/10.1006/jsbi.2000.4267
- Primary Citation Related Structures:
1BJB, 1BJC - PubMed Abstract:
NMRsolution structures are reported for two mutants (K16E, K16F) of the soluble amyloid beta peptide Abeta(1-28). The structural effects of these mutations of a positively charged residue to anionic and hydrophobic residues at the alpha-secretase cleavage site (Lys16-Leu17) were examined in the membrane-simulating solvent aqueous SDS micelles. Overall the three-dimensional structures were similar to that for the native Abeta(1-28) sequence in that they contained an unstructured N-terminus and a helical C-terminus. These structural elements are similar to those seen in the corresponding regions of full-length Abeta peptides Abeta(1-40) and Abeta(1-42), showing that the shorter peptides are valid model systems. The K16E mutation, which might be expected to stabilize the macrodipole of the helix, slightly increased the helix length (residues 13-24) relative to the K16F mutation, which shortened the helix to between residues 16 and 24. The observed sequence-dependent control over conformation in this region provides an insight into possible conformational switching roles of mutations in the amyloid precursor protein from which Abeta peptides are derived. In addition, if conformational transitions from helix to random coil to sheet precede aggregation of Abeta peptides in vivo, as they do in vitro, the conformation-inducing effects of mutations at Lys16 may also influence aggregation and fibril formation.
- Institute for Molecular Bioscience, The University of Queensland, Brisbane, QLD 4072, Australia.
Organizational Affiliation: