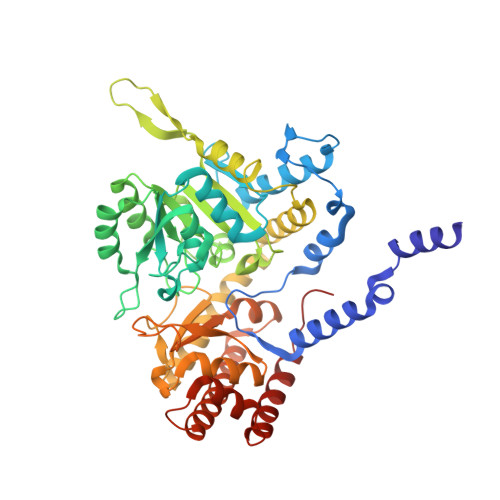
The crystal structure of human cytosolic serine hydroxymethyltransferase: a target for cancer chemotherapy
Renwick, S.B., Snell, K., Baumann, U.(1998) Structure 6: 1105-1116
- PubMed: 9753690 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00112-9
- Primary Citation Related Structures:
1BJ4 - PubMed Abstract:
Serine hydroxymethyltransferase (SHMT) is a ubiquitous enzyme found in all prokaryotes and eukaryotes. As an enzyme of the thymidylate synthase metabolic cycle, SHMT catalyses the retro-aldol cleavage of serine to glycine, with the resulting hydroxymethyl group being transferred to tetrahydrofolate to form 5, 10-methylene-tetrahydrofolate. The latter is the major source of one-carbon units in metabolism. Elevated SHMT activity has been shown to be coupled to the increased demand for DNA synthesis in rapidly proliferating cells, particularly tumour cells. Consequently, the central role of SHMT in nucleotide biosynthesis makes it an attractive target for cancer chemotherapy. We have solved the crystal structure of human cytosolic SHMT by multiple isomorphous replacement to 2.65 A resolution. The monomer has a fold typical for alpha class pyridoxal 5'-phosphate (PLP) dependent enzymes. The tetramer association is best described as a 'dimer of dimers' where residues from both subunits of one 'tight' dimer contribute to the active site. The crystal structure shows the evolutionary relationship between SHMT and other alpha class PLP-dependent enzymes, as the fold is highly conserved. Many of the results of site-directed mutagenesis studies can easily be rationalised or re-interpreted in light of the structure presented here. For example, His 151 is not the catalytic base, contrary to the findings of others. A mechanism for the cleavage of serine to glycine and formaldehyde is proposed.
- Section of Structural Biology Institute of Cancer Research University of London Cotswold Road, Sutton, Surrey, SM2 5NG, Celltech plc 216 Bath Road, Slough, Berkshire, SL1 4EN, UK.
Organizational Affiliation: