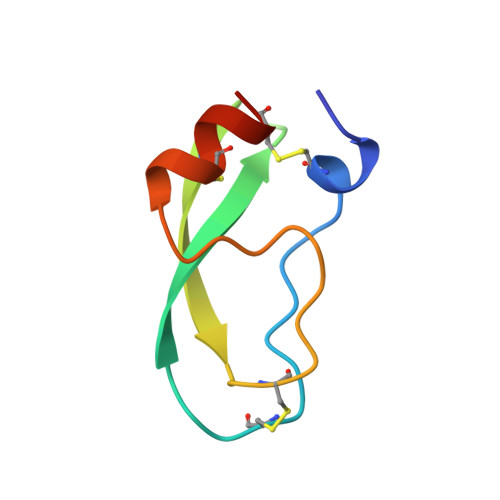
The decameric structure of bovine pancreatic trypsin inhibitor (BPTI) crystallized from thiocyanate at 2.7 A resolution.
Hamiaux, C., Prange, T., Ries-Kautt, M., Ducruix, A., Lafont, S., Astier, J.P., Veesler, S.(1999) Acta Crystallogr D Biol Crystallogr 55: 103-113
- PubMed: 10089400 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444998008725
- Primary Citation Related Structures:
1BHC - PubMed Abstract:
The structure of a monoclinic form of bovine pancreatic trypsin inhibitor (BPTI) crystallized from a thiocyanate solution has been determined and refined at 2.7 A resolution. The space group is P21 with a = 71.56, b = 73.83, c = 64.47 A, beta = 93.9 degrees and Z = 20. The ten independent molecules were located by a multi-body molecular-replacement search as developed in the AMoRe program, starting from a single monomer model (PDB code: 6PTI). The molecular arrangement of the subunits is a decamer resulting from the combination of two orthogonal fivefold and twofold non-crystallographic axes. This builds a globular micelle-like particle which minimizes hydrophobic interactions with the solvent. The refinement was conducted with non-crystallographic symmetry constraints up to a final residual of R = 0.20 (Rfree= 0.26). The root-mean-square deviations from ideal geometry were 0.015 A and 1.6 degrees on bond distances and bond angles, respectively. Several sites for thiocyanate ions were analyzed.
- Laboratoire pour l'Utilisation du Rayonnement Electromagnétique (LURE), Bâtiment 209d, Université Paris-Sud, 91405 Orsay CEDEX, France.
Organizational Affiliation: