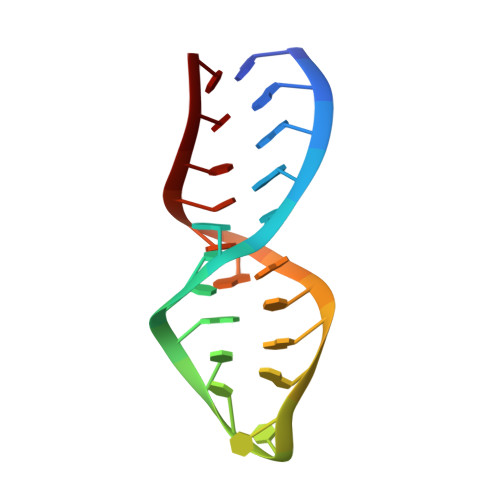
NMR structure determination of the binding site for ribosomal protein S8 from Escherichia coli 16 S rRNA.
Kalurachchi, K., Nikonowicz, E.P.(1998) J Mol Biology 280: 639-654
- PubMed: 9677294
- DOI: https://doi.org/10.1006/jmbi.1998.1915
- Primary Citation Related Structures:
1BGZ - PubMed Abstract:
Many cellular processes involve the preferential interaction of an RNA molecule with a specific protein. A detailed analysis of the individual protein and RNA components of these interactions can provide unique insights into the structural features important for protein-RNA recognition. Ribosomal protein S8 of Escherichia coli plays a key role in 30 S ribosomal subunit assembly through its interaction with 16 S rRNA. The binding site for protein S8 comprises a portion of helix 21, nucleotides G588 to G604 and C634 to C651. This region forms a base-paired helix that is interrupted by a non-Watson-Crick segment composed of nine phylogenetically conserved nucleotides. We have investigated the detailed structure of the conserved segment and the interaction of this region with metal ions using NMR spectroscopy. Twenty-four of the 40 calculated structures converged to similar conformations and were grouped into two families. The main difference between the families is the orientation of the base of U641. The rms deviation between the heavy-atoms of the ten lowest-energy structures is 1.24 A. The orientations of the G597.C643 base-pair and A595.(A596.U644) base-triple within the conserved core have been defined and appear to extend the proximal segment of helix 21 into the phylogenetically conserved core. The base of A642 terminates this helix by stacking against C643 and the base of U641 forms hydrogen bonds with core nucleotides. The conserved core also contains a Mg2+-binding site that promotes stabilization of the secondary and tertiary structure elements of the core. A model for the interaction of S8 with its RNA-binding site is proposed.
- Department of Biochemistry and Cell Biology, Rice University, Houston, TX 77005, USA.
Organizational Affiliation: