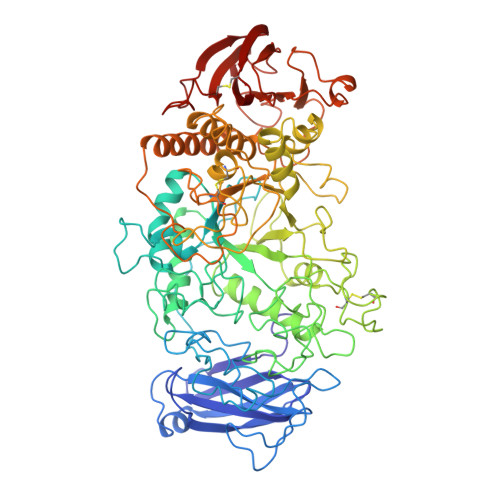
Three-dimensional structure of Pseudomonas isoamylase at 2.2 A resolution.
Katsuya, Y., Mezaki, Y., Kubota, M., Matsuura, Y.(1998) J Mol Biology 281: 885-897
- PubMed: 9719642
- DOI: https://doi.org/10.1006/jmbi.1998.1992
- Primary Citation Related Structures:
1BF2 - PubMed Abstract:
The three-dimensional structure of isoamylase from Pseudomonas amyloderamosa, which hydrolyzes alpha-1,6-glucosidic linkages of amylopectin and glycogen, has been determined by X-ray structure analysis. The enzyme has 750 amino acid residues and a molecular mass of 80 kDa, and it can be crystallized from ammonium sulfate solution. The structure was elucidated by the multiple isomorphous replacement method and refined at 2.2 A resolution, resulting in a final R-factor of 0.161 for significant reflections with a root-mean-square deviation from ideality in bond lengths of 0.009 A. The analysis revealed that in the N-terminal region, isoamylase has a novel extra domain that we call domain N, whose three-dimensional structure has not so far been reported. It has a (beta/alpha)8-barrel-type supersecondary structure in the catalytic domain common to the alpha-amylase family enzymes, though the barrel is incomplete, with a deletion of an alpha-helix between the fifth and sixth beta-strands. A long excursed region is present between the third beta-strand and the third alpha-helix of the barrel but, in contrast to the so-called domain B that has been identified in the other enzymes of alpha-amylase family, it cannot be considered to be an independent domain, because this loop forms a globular cluster together with the loop between the fourth beta-strand and the fourth alpha-helix. Isoamylase contains a bound calcium ion, but this is not in the same position as the conserved calcium ion that has been reported in other alpha-amylase family enzymes.
- Hyogo Prefectural Institute of Industrial Research, Suma-ku, Kobe, 654-0037, Japan. ykatsuya@hyogo-kp.go.jp
Organizational Affiliation: