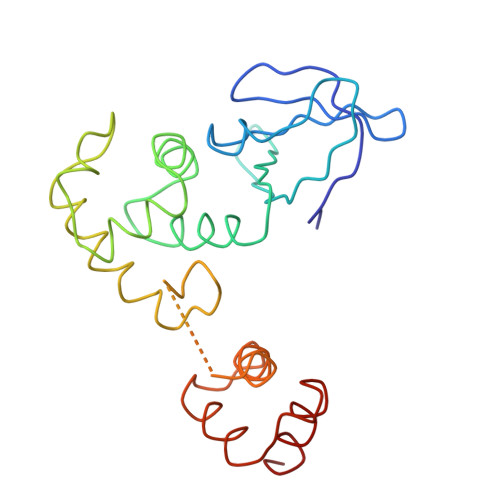
Crystal structure of E.coli RuvA with bound DNA Holliday junction at 6 A resolution.
Hargreaves, D., Rice, D.W., Sedelnikova, S.E., Artymiuk, P.J., Lloyd, R.G., Rafferty, J.B.(1998) Nat Struct Biol 5: 441-446
- PubMed: 9628481 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0698-441
- Primary Citation Related Structures:
1BDX - PubMed Abstract:
Here we present the crystal structure of the Escherichia coli protein RuvA bound to a key DNA intermediate in recombination, the Holliday junction. The structure, solved by isomorphous replacement and density modification at 6 A resolution, reveals the molecular architecture at the heart of the branch migration and resolution reactions required to process Holliday intermediates into recombinant DNA molecules. It also reveals directly for the first time the structure of the Holliday junction. A single RuvA tetramer is bound to one face of a junction whose four DNA duplex arms are arranged in an open and essentially four-fold symmetric conformation. Protein-DNA contacts are mediated by two copies of a helix-hairpin-helix motif per RuvA subunit that contact the phosphate backbone in a very similar manner. The open structure of the junction stabilized by RuvA binding exposes a DNA surface that could be bound by the RuvC endonuclease to promote resolution.
- Krebs Institute, Department of Molecular Biology and Biotechnology, University of Sheffield, Western Bank, UK.
Organizational Affiliation: