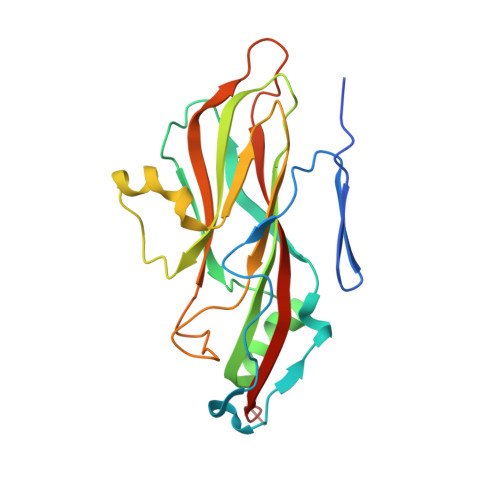
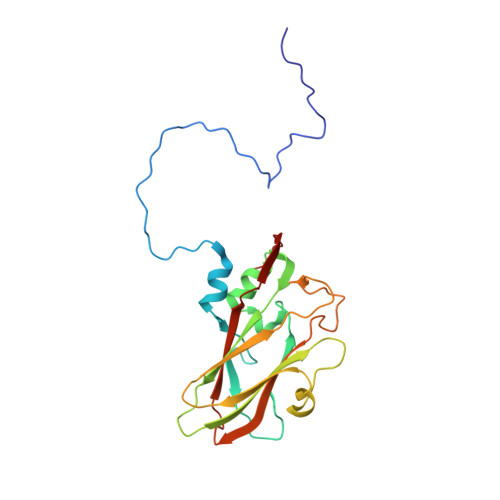
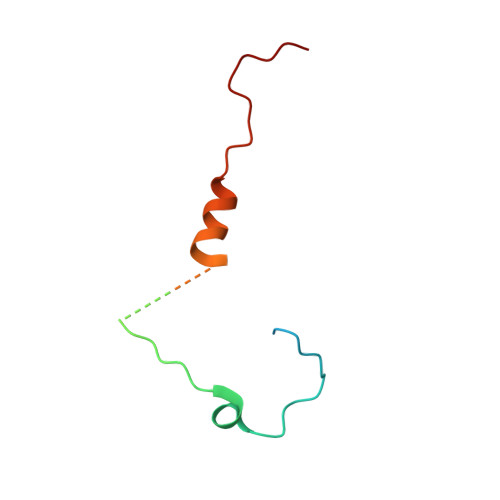
Methods used in the structure determination of foot-and-mouth disease virus.
Fry, E., Acharya, R., Stuart, D.(1993) Acta Crystallogr A 49: 45-55
- PubMed: 8382928
- DOI: https://doi.org/10.1107/s0108767392005737
- Primary Citation of Related Structures:
1BBT - PubMed Abstract:
The structure of foot-and-mouth disease virus (FMDV) strain O1 BFS 1860 has been determined to 2.9 A resolution using the molecular-replacement method [Acharya, Fry, Stuart, Fox, Rowlands & Brown (1989). Nature (London), 337, 709-716]. Crystals of the virus with average dimensions 0.12 x 0.06 x 0.12 mm belong to space group I23, a = 345 A with 1/12 of the icosahedral particle per asymmetric unit giving fivefold noncrystallographic redundancy. Oscillation diffraction photographs were collected at the SERC Synchrotron Radiation Source at Daresbury in accordance with strict disease security regulations. The ambiguity in particle orientation was resolved using a self-rotation function and starting estimates of the phases to 8 A were derived from the known structures of two picornaviruses similarly oriented in the I23 unit cell. The phases were refined and extended using iterative averaging and solvent flattening with the implementation of a simple automatic envelope-determination procedure to increase the phasing power available.
- Laboratory of Molecular Biophysics, Oxford, England.
Organizational Affiliation: