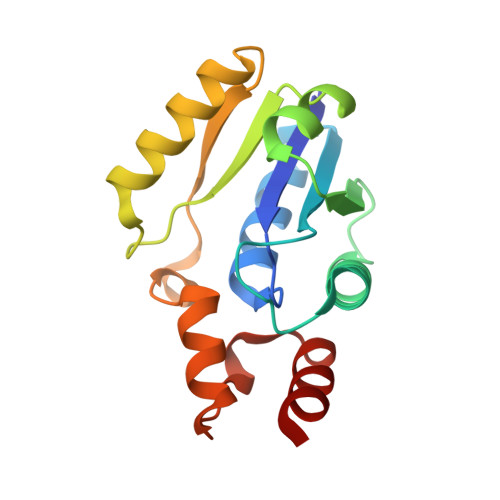
The crystal structure of a novel bacterial adenylyltransferase reveals half of sites reactivity.
Izard, T., Geerlof, A.(1999) EMBO J 18: 2021-2030
- PubMed: 10205156
- DOI: https://doi.org/10.1093/emboj/18.8.2021
- Primary Citation of Related Structures:
1B6T - PubMed Abstract:
Phosphopantetheine adenylyltransferase (PPAT) is an essential enzyme in bacteria that catalyses a rate-limiting step in coenzyme A (CoA) biosynthesis, by transferring an adenylyl group from ATP to 4'-phosphopantetheine, yielding dephospho-CoA (dPCoA). Each phosphopantetheine adenylyltransferase (PPAT) subunit displays a dinucleotide-binding fold that is structurally similar to that in class I aminoacyl-tRNA synthetases. Superposition of bound adenylyl moieties from dPCoA in PPAT and ATP in aminoacyl-tRNA synthetases suggests nucleophilic attack by the 4'-phosphopantetheine on the alpha-phosphate of ATP. The proposed catalytic mechanism implicates transition state stabilization by PPAT without involving functional groups of the enzyme in a chemical sense in the reaction. The crystal structure of the enzyme from Escherichia coli in complex with dPCoA shows that binding at one site causes a vice-like movement of active site residues lining the active site surface. The mode of enzyme product formation is highly concerted, with only one trimer of the PPAT hexamer showing evidence of dPCoA binding. The homologous active site attachment of ATP and the structural distribution of predicted sequence-binding motifs in PPAT classify the enzyme as belonging to the nucleotidyltransferase superfamily.
- Department of Biochemistry, University of Leicester, Leicester LE1 7RH.ti3@le.ac.uk
Organizational Affiliation: