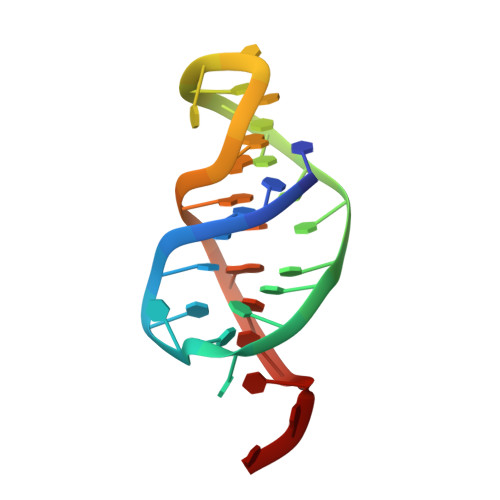
Structure and mechanism of formation of the H-y5 isomer of an intramolecular DNA triple helix.
van Dongen, M.J., Doreleijers, J.F., van der Marel, G.A., van Boom, J.H., Hilbers, C.W., Wijmenga, S.S.(1999) Nat Struct Biol 6: 854-859
- PubMed: 10467098 Search on PubMed
- DOI: https://doi.org/10.1038/12313
- Primary Citation Related Structures:
1B4Y - PubMed Abstract:
H-DNA, thought to play a regulatory role in transcription, exists in two isomeric forms, H-y3 and H-y5. We present the first solution structure of a DNA fragment representing the H-y5 fold. The structure shows the H-y5 triple helix, and for the first time how in an H-DNA isomer the purine strand extension interacts with the triplex loop. It gives direct insight into the mechanism of H-DNA formation, and explains a host of biochemical and biophysical data on the relative stability of the H-DNA isomers. In addition, the observed interaction of the purine strand extension and the triplex loop provides new clues to the design of clamp-type triple helix-forming oligonucleotides.
- NSR Centre for Molecular Structure, Design, and Synthesis, Laboratory of Biophysical Chemistry, University of Nijmegen, The Netherlands.
Organizational Affiliation: