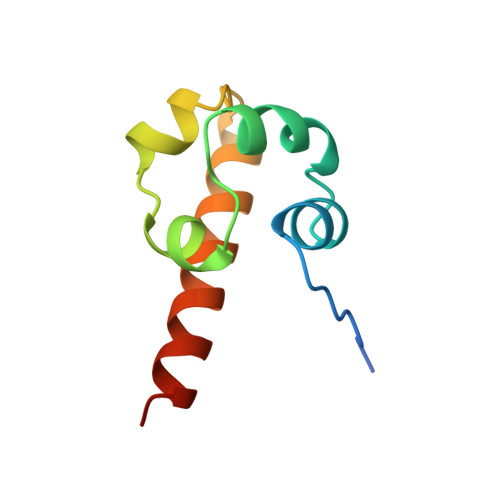
Oligomeric structure of the human EphB2 receptor SAM domain.
Thanos, C.D., Goodwill, K.E., Bowie, J.U.(1999) Science 283: 833-836
- PubMed: 9933164 Search on PubMed
- DOI: https://doi.org/10.1126/science.283.5403.833
- Primary Citation Related Structures:
1B4F - PubMed Abstract:
The sterile alpha motif (SAM) domain is a protein interaction module that is present in diverse signal-transducing proteins. SAM domains are known to form homo- and hetero-oligomers. The crystal structure of the SAM domain from an Eph receptor tyrosine kinase, EphB2, reveals two large interfaces. In one interface, adjacent monomers exchange amino-terminal peptides that insert into a hydrophobic groove on each neighbor. A second interface is composed of the carboxyl-terminal helix and a nearby loop. A possible oligomer, constructed from a combination of these binding modes, may provide a platform for the formation of larger protein complexes.
- UCLA-DOE Laboratory of Structural Biology and Molecular Medicine and Department of Chemistry and Biochemistry, University of California, Los Angeles, CA 90095, USA.
Organizational Affiliation: