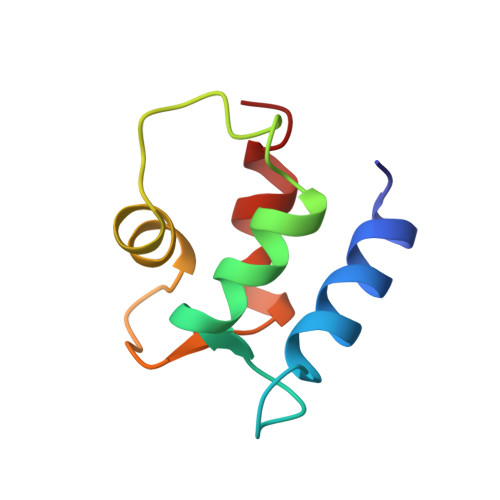
Protein solution structure calculations in solution: solvated molecular dynamics refinement of calbindin D9k.
Kordel, J., Pearlman, D.A., Chazin, W.J.(1997) J Biomol NMR 10: 231-243
- PubMed: 9390401 Search on PubMed
- DOI: https://doi.org/10.1023/a:1018383102870
- Primary Citation Related Structures:
1B1G - PubMed Abstract:
The three-dimensional solution structures of proteins determined with NMR-derived constraints are almost always calculated in vacuo. The solution structure of (Ca2+)2-calbindin D9k has been redetermined by new restrained molecular dynamics (MD) calculations that include Ca2+ ions and explicit solvent molecules. Four parallel sets of MD refinements were run to provide accurate comparisons of structures produced in vacuo, in vacuo with Ca2+ ions, and with two different protocols in a solvent bath with Ca2+ ions. The structural ensembles were analyzed in terms of structural definition, molecular energies, packing density, solvent-accessible surface, hydrogen bonds, and the coordination of calcium ions in the two binding loops. Refinement including Ca2+ ions and explicit solvent results in significant improvements in the precision and accuracy of the structure, particularly in the binding loops. These results are consistent with results previously obtained in free MD simulations of proteins in solution and show that the rMD refined NMR-derived solution structures of proteins, especially metalloproteins, can be significantly improved by these strategies.
- Pharmacia & Upjohn, Stockholm, Sweden.
Organizational Affiliation: