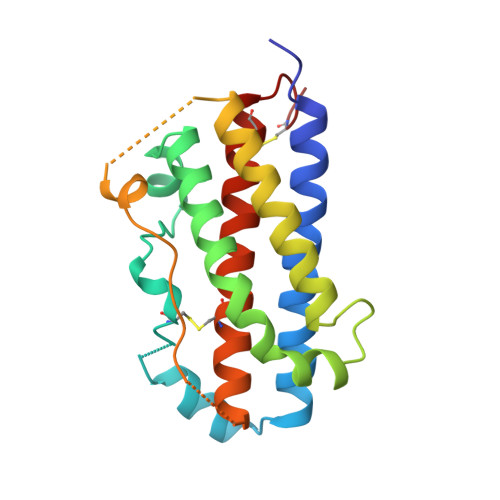
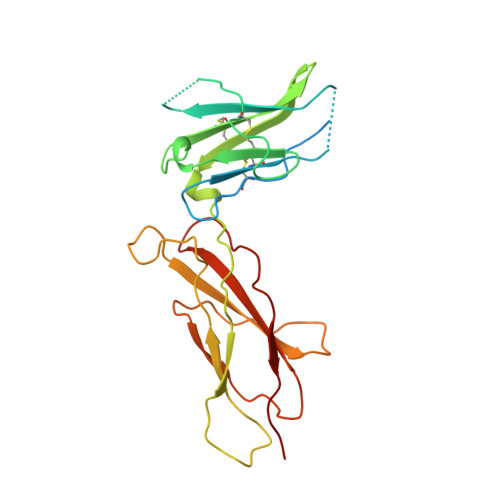
Structural plasticity in a remodeled protein-protein interface.
Atwell, S., Ultsch, M., De Vos, A.M., Wells, J.A.(1997) Science 278: 1125-1128
- PubMed: 9353194
- DOI: https://doi.org/10.1126/science.278.5340.1125
- Primary Citation of Related Structures:
1AXI - PubMed Abstract:
Remodeling of the interface between human growth hormone (hGH) and the extracellular domain of its receptor was studied by deleting a critical tryptophan residue (at position 104) in the receptor, creating a large cavity, and selecting a pentamutant of hGH by phage display that fills the cavity and largely restores binding affinity. A 2.1 A resolution x-ray structure of the mutant complex showed that the receptor cavity was filled by selected hydrophobic mutations of hGH. Large structural rearrangements occurred in the interface at sites that were distant from the mutations. Such plasticity may be a means for protein-protein interfaces to adapt to mutations as they coevolve.
- Department of Protein Engineering, Genentech, Incorporated, 460 Point San Bruno Boulevard, South San Francisco, CA 94080, USA.
Organizational Affiliation: