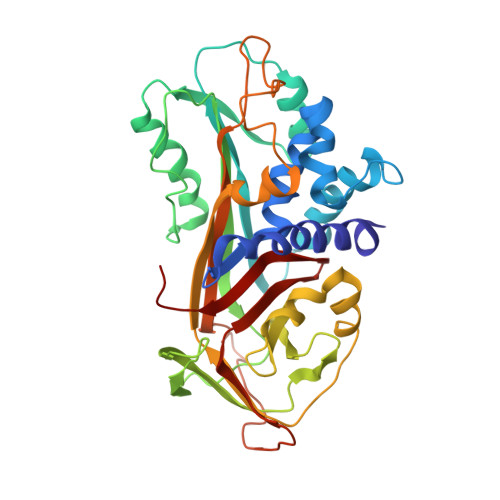
The native strains in the hydrophobic core and flexible reactive loop of a serine protease inhibitor: crystal structure of an uncleaved alpha1-antitrypsin at 2.7 A.
Ryu, S.E., Choi, H.J., Kwon, K.S., Lee, K.N., Yu, M.H.(1996) Structure 4: 1181-1192
- PubMed: 8939743 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(96)00126-8
- Primary Citation Related Structures:
1ATU - PubMed Abstract:
The protein alpha1-antitrypsin is a prototype member of the serpin (serine protease inhibitor) family and is known to inhibit the activity of neutrophil elastase in the lower respiratory tract. Members of this family undergo a large structural rearrangement upon binding to a target protease, involving cleavage of the reactive-site loop. This loop is then inserted into the main body of the enzyme following the opening of a central beta sheet, leading to stabilization of the structure. Random mutageneses of alpha1-antitrypsin identified various mutations that stabilize the native structure and retard the insertion of the reactive-site loop. Structural studies of these mutations may reveal the mechanism of the conformational change. We have determined the three-dimensional structure of an uncleaved alpha1-antitrypsin with seven such stabilizing mutations (hepta alpha1-antitrypsin) at 2.7 A resolution. From the comparison of the structure with other serpin structures, we found that hepta alpha1-antitrypsin is stabilized due to the release of various strains that exist in native wild type alpha1-antitrypsin, including unfavorable hydrophobic interactions in the central hydrophobic core. The reactive-site loop of hepta alpha1-antitrypsin is an extended strand, different from that of the previously determined structure of another uncleaved alpha1-antitrypsin, and indicates the inherent flexibility of the loop. The present structural study suggests that the uncleaved alpha1-antitrypsin has many folding defects which can be improved by mutations. These folding defects seem to be utilized in a coordinated fashion in the regulation of the conformational switch of alpha1-antitrypsin. Some of the defects, represented by the Phe51 region and possibly the Met374 and the Thr59 regions, are part of the sheet-opening mechanism.
- Protein Engineering Research Division, Korea Research Institute of Bioscience and Biotechnology, KIST, Yusong, Taejon, South Korea. ryuse@mcr1.geri.re.kr
Organizational Affiliation: