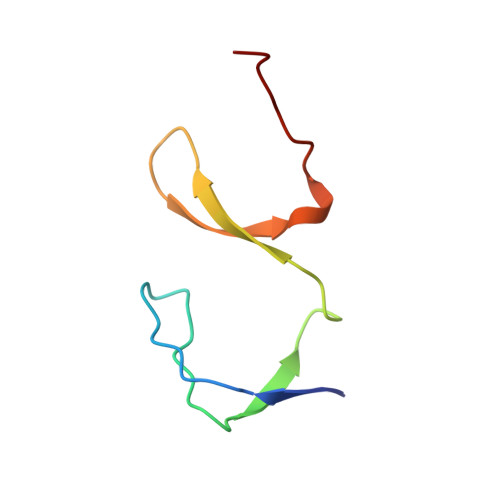
The SH3 domain of Eps8 exists as a novel intertwined dimer.
Kishan, K.V., Scita, G., Wong, W.T., Di Fiore, P.P., Newcomer, M.E.(1997) Nat Struct Biol 4: 739-743
- PubMed: 9303002 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0997-739
- Primary Citation Related Structures:
1AOJ - PubMed Abstract:
SH3 domains are structurally well-characterized as monomeric modular units of protein structure that mediate protein-protein recognition in numerous signal transduction proteins. The X-ray crystallographic structure of the Eps8 SH3 domain reveals a novel variation of the canonical SH3 fold: the SH3 domain from Eps8 is a dimer formed by strand interchange. In addition, co-immunoprecipitation experiments show that intact Eps8 is multimeric in vivo. Hence, the SH3 domain of Eps8 may represent a dimerization motif.
- Department of Biochemistry, Vanderbilt University School of Medicine, Nashville, TN 37232-0146, USA.
Organizational Affiliation: