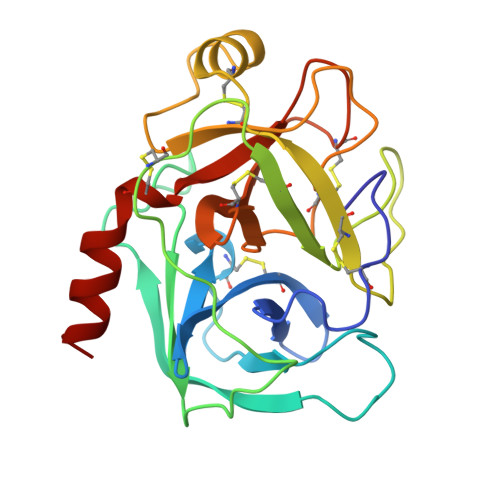
Structure of an engineered, metal-actuated switch in trypsin.
McGrath, M.E., Haymore, B.L., Summers, N.L., Craik, C.S., Fletterick, R.J.(1993) Biochemistry 32: 1914-1919
- PubMed: 8448149 Search on PubMed
- DOI: https://doi.org/10.1021/bi00059a005
- Primary Citation Related Structures:
1AND, 1ANE - PubMed Abstract:
The X-ray crystal structure of the copper complex of the rat trypsin mutant Arg96 to His96 (trypsin R96H) has been determined in order to ascertain the nature of the engineered metal-binding site and to understand the structural basis for the metal-induced enzymatic inhibition. In the structure, the catalytically essential His57 residue is reoriented out of the active-site pocket and forms a chelating, metal-binding site with residue His96. The copper is bound to the N epsilon 2 atoms of both histidine residues with Cu-N epsilon 2 = 2.2 A and N epsilon 2-Cu-N epsilon 2 = 89 degrees. The metal is clearly bound to a third ligand leading to a distorted square planar geometry at Cu. The X-ray results do not unambiguously yield the identity of this third ligand, but chemical data suggest that it is a deprotonated, chelating Tris molecule which was used as a carrier to solubilize the copper in alkaline solution (pH 8.0). Upon reorientation of His57, a unique water molecule moves into the active site and engages in hydrogen-bonding with Asp102-O delta 2 and His57-N delta 1. Except for small movements of the peptide backbone near His96, the remainder of the trypsin molecule is isostructural with the native enzyme. These data support the notion that the effective inhibition of catalytic activity by metal ions observed in trypsin R96H is indeed caused by a specific and reversible reorganization of the active site in the enzyme.
- Department of Biochemistry and Biophysics, University of California, San Francisco 94143.
Organizational Affiliation: