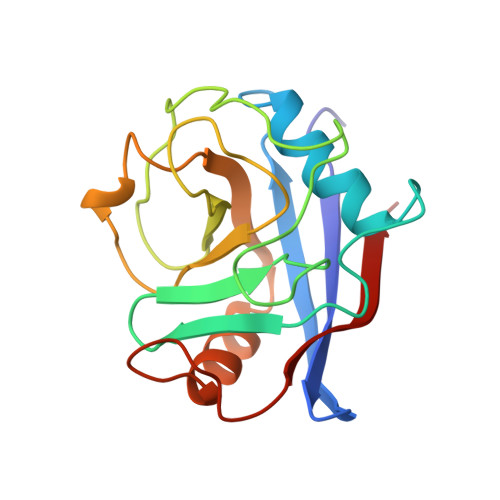
Crystal structure of human cyclophilin A bound to the amino-terminal domain of HIV-1 capsid.
Gamble, T.R., Vajdos, F.F., Yoo, S., Worthylake, D.K., Houseweart, M., Sundquist, W.I., Hill, C.P.(1996) Cell 87: 1285-1294
- PubMed: 8980234 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81823-1
- Primary Citation Related Structures:
1AK4 - PubMed Abstract:
The HIV-1 capsid protein forms the conical core structure at the center of the mature virion. Capsid also binds the human peptidyl prolyl isomerase, cyclophilin A, thereby packaging the enzyme into the virion. Cyclophilin A subsequently performs an essential function in HIV-1 replication, possibly helping to disassemble the capsid core upon infection. We report the 2.36 A crystal structure of the N-terminal domain of HIV-1 capsid (residues 1-151) in complex with human cyclophilin A. A single exposed capsid loop (residues 85-93) binds in the enzyme's active site, and Pro-90 adopts an unprecedented trans conformation. The structure suggests how cyclophilin A can act as a sequence-specific binding protein and a nonspecific prolyl isomerase. In the crystal lattice, capsid molecules assemble into continuous planar strips. Side by side association of these strips may allow capsid to form the surface of the viral core. Cyclophilin A could then function by weakening the association between capsid strips, thereby promoting disassembly of the viral core.
- Biochemistry Department, University of Utah, Salt Lake City 84103, USA.
Organizational Affiliation: