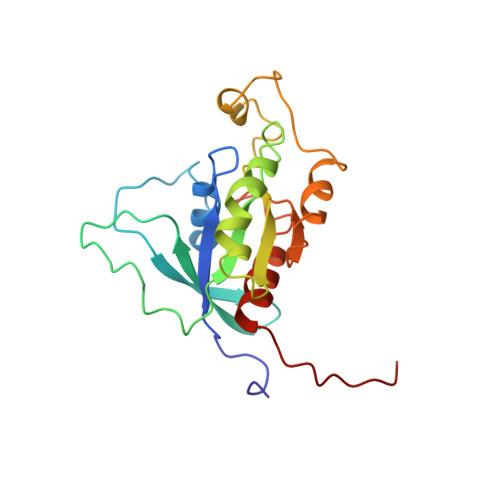
Definition of the switch surface in the solution structure of Cdc42Hs.
Feltham, J.L., Dotsch, V., Raza, S., Manor, D., Cerione, R.A., Sutcliffe, M.J., Wagner, G., Oswald, R.E.(1997) Biochemistry 36: 8755-8766
- PubMed: 9220962 Search on PubMed
- DOI: https://doi.org/10.1021/bi970694x
- Primary Citation Related Structures:
1AJE - PubMed Abstract:
Proteins of the rho subfamily of ras GTPases have been shown to be crucial components of pathways leading to cell growth and the establishment of cell polarity and mobility. Presented here is the solution structure of one such protein, Cdc42Hs, which provides insight into the structural basis for specificity of interactions between this protein and its effector and regulatory proteins. Standard heteronuclear NMR methods were used to assign the protein, and approximately 2100 distance and dihedral angle constraints were used to calculate a set of 20 structures using a combination of distance geometry and simulated annealing refinement. These structures show overall similarity to those of other GTP-binding proteins, with some exceptions. The regions corresponding to switch I and switch II in H-ras are disordered, and no evidence was found for an alpha-helix in switch II. The 13-residue insertion, which is only present in rho-subtype proteins and has been shown to be an important mediator of binding of regulatory and target proteins, forms a compact structure containing a short helix lying adjacent to the beta4-alpha3 loop. The insert forms one edge of a "switch surface" and, unexpectedly, does not change conformation upon activation of the protein by the exchange of GTP analogs for GDP. These studies indicate the insert region forms a stable invariant "footrest" for docking of regulatory and effector proteins.
- Department of Pharmacology, College of Veterinary Medicine, Cornell University, Ithaca, New York 14853, USA.
Organizational Affiliation: