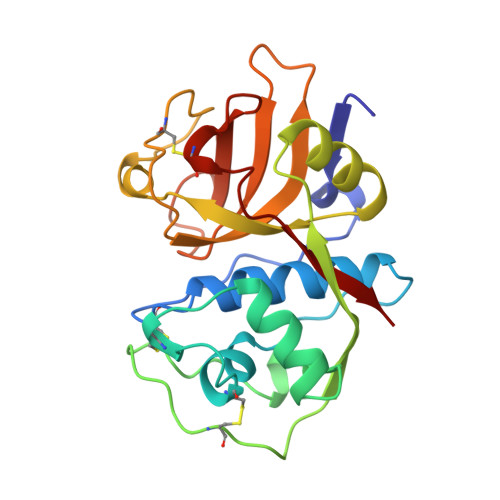
Structural determinants of specificity in the cysteine protease cruzain.
Gillmor, S.A., Craik, C.S., Fletterick, R.J.(1997) Protein Sci 6: 1603-1611
- PubMed: 9260273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560060801
- Primary Citation Related Structures:
1AIM, 2AIM - PubMed Abstract:
The structure of cruzain, an essential protease from the parasite Trypanosoma cruzi, was determined by X-ray crystallography bound to two different covalent inhibitors. The cruzain S2 specificity pocket is able to productively bind both arginine and phenylalanine residues. The structures of cruzain bound to benzoyl-Arg-Ala-fluoromethyl ketone and benzoyl-Tyr-Ala-fluoromethyl ketone at 2.2 and 2.1 A, respectively, show a pH-dependent specificity switch. Glu 205 adjusts to restructure the S2 specificity pocket, conferring right binding to both hydrophobic and basic residues. Kinetic analysis of activated peptide substrates shows that substrates placing hydrophobic residues in the specificity pocket are cleaved at a broader pH range than hydrophilic substrates. These results demonstrate how cruzain binds both basic and hydrophobic residues and could be important for in vivo regulation of cruzain activity.
- Graduate Group in Biophysics, University of California, San Francisco 94143-0448, USA.
Organizational Affiliation: