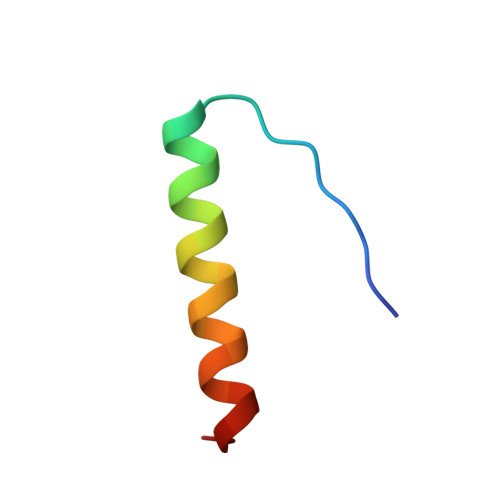
Crystallization and structure solution of p53 (residues 326-356) by molecular replacement using an NMR model as template.
Mittl, P.R., Chene, P., Grutter, M.G.(1998) Acta Crystallogr D Biol Crystallogr 54: 86-89
- PubMed: 9761820 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444997006550
- Primary Citation Related Structures:
1AIE - PubMed Abstract:
The molecular replacement method is a powerful technique for crystal structure solution but the use of NMR structures as templates often causes problems. In this work the NMR structure of the p53 tetramerization domain has been used to solve the crystal structure by molecular replacement. Since the rotation- and translation-functions were not sufficiently clear, additional information about the symmetry of the crystal and the protein complex was used to identify correct solutions. The three-dimensional structure of residues 326-356 was subsequently refined to a final R factor of 19.1% at 1.5 A resolution.
- Core Drug Discovery Technologies, Ciba-Geigy AG, CH-4002 Basel, Switzerland. mittl@fmi.ch
Organizational Affiliation: