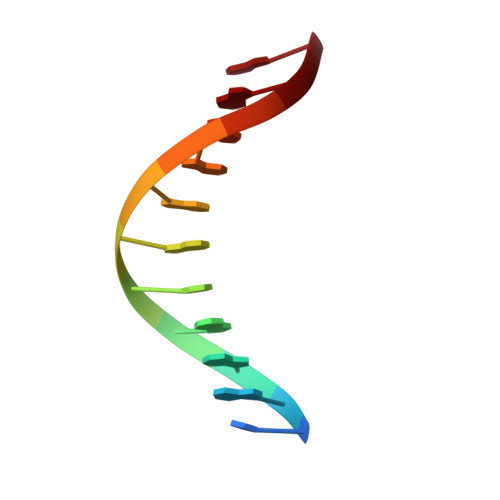
Solution structure of an oligodeoxynucleotide containing the human n-ras codon 61 sequence refined from 1H NMR using molecular dynamics restrained by nuclear Overhauser effects.
Feng, B., Stone, M.P.(1995) Chem Res Toxicol 8: 821-832
- PubMed: 7492731
- DOI: https://doi.org/10.1021/tx00048a002
- Primary Citation Related Structures:
1AGH - PubMed Abstract:
The solution structure of the ras61 oligodeoxynucleotide duplex d(CGGACAAGAAG). d(CTTCTTGTCCG), which consists of codons 60, 61 (underlined), and 62 of the human n-ras protooncogene, was refined from 1H NMR data. The sequence contains a run of purines in the coding strand, with one R-Y step, A4.T19-->C5.G18, and one Y-R step, C5.G18-->A6.T17 (excluding the 5'-terminal base pair). The NMR data were consistent with a B-like helix as judged by characteristic internucleotide NOEs. The NOE intensities between purine H8 and purine anomeric protons were small as compared to the intensities between cytosine H5 and H6 protons, indicative of glycosyl torsion angles in the anti range. Cross-peaks were observed between purine H8 and pyrimidine H5 and CH3 protons on adjacent bases in the direction of purine (5'-->3') pyrimidine, but not in the direction pyrimidine (5'-->3') purine. Watson-Crick hydrogen bonding was intact and enabled the assignment of the exchangeable protons. A total of 226 experimental distance restraints were obtained. A restrained molecular dynamics and simulated annealing approach was utilized in the refinement. The data for 5 emergent molecular dynamics (MD) structures calculated from a B-form starting structure and 5 emergent MD structures calculated from an A-form starting structure refined to an average pairwise root-mean-square (rms) difference of 1.2 A, with maximum pairwise rmsd of 1.7 A. The accuracy of the emergent structures was assessed by complete relaxation matrix back-calculation. The sixth root residual index of 9.4 x 10(-2) was measured between the refined structures and the NOE data, suggesting that the former were in reasonable agreement with the data. The refined structures revealed an increased roll angle of 7 degrees in the codon 61 sequence at base step C5.G18-->A6.T17, which relieved the purine-purine clash in the minor groove, and in turn relieved the purine-purine clash in major groove between A4.T19 and C5.G18. A 3.7 A rise between C5.G18 and A6.T17 was calculated, which assisted in relieving the purine-purine clash. The local variations in the B-like conformation did not confer large structural alterations upon the ras61 sequence, but could be important in modulating the reactivity of the first as compared to the second adenine in codon 61.
- Department of Chemistry, Vanderbilt University, Nashville, Tennessee 37235, USA.
Organizational Affiliation: