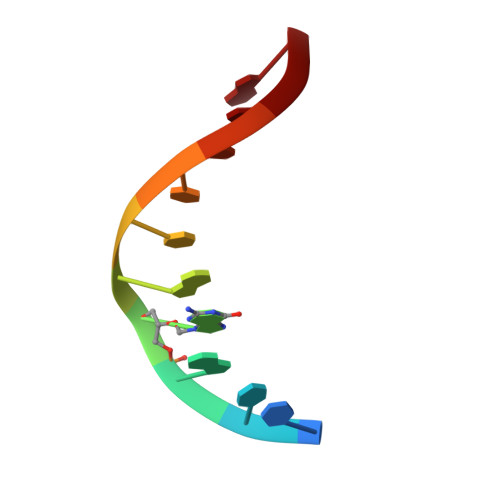
Solution structure of a DNA decamer containing the antiviral drug ganciclovir: combined use of NMR, restrained molecular dynamics, and full relaxation matrix refinement.
Foti, M., Marshalko, S., Schurter, E., Kumar, S., Beardsley, G.P., Schweitzer, B.I.(1997) Biochemistry 36: 5336-5345
- PubMed: 9154915
- DOI: https://doi.org/10.1021/bi962604e
- Primary Citation Related Structures:
1AC9 - PubMed Abstract:
The nucleoside analog 9-[(1,3-dihydroxy-2-propoxy)methyl]guanine (ganciclovir, DHPG) is an antiviral drug that is used in the treatment of a variety of herpes viruses in immunocompromised patients and in a gene therapy protocol that has shown promising activity for the treatment of cancer. To probe the structural effects of ganciclovir when incorporated into DNA, we determined and compared the solution structure of a modified ganciclovir-containing decamer duplex [d(CTG)(ganciclovir)d(ATCCAG)]2 and a control duplex d[(CTGGATCCAG)]2 using nuclear magnetic resonance techniques. 1H and 31P resonances in both duplexes were assigned using a combination of 2-D 1H and 31P NMR experiments. Proton-proton distances determined from NOESY data and dihedral angles determined from DQF-COSY data were used in restrained molecular dynamics simulations starting from canonical A- and B-form DNA models. Both the control and ganciclovir sets of simulations converged to B-type structures. These structures were subjected to full relaxation matrix refinement to produce final structures that were in excellent agreement with the observed NOE intensities. Examination of the final ganciclovir-containing structures reveals that the base of the ganciclovir residue is hydrogen bonded to its complementary dC and is stacked in the helix; in fact, the base of ganciclovir exhibits increased stacking with the 5' base relative to the control. Interestingly, some of the most significant distortions in the structures occur 3' to the lesion site, including a noticeable kink in the sugar-phosphate backbone at this position. Further examination reveals that the backbone conformation, sugar pucker, and glycosidic torsion angle of the residue 3' to the lesion site all indicate an A-type conformation at this position. A possible correlation of these structural findings with results obtained from earlier biochemical studies will be discussed.
- Walt Disney Memorial Cancer Institute at Florida Hospital, Orlando 32826, USA.
Organizational Affiliation: