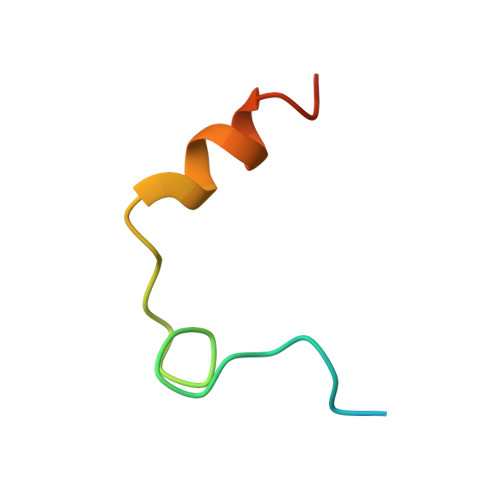
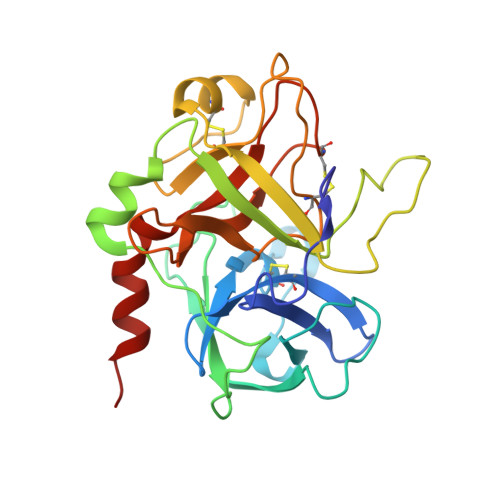
Structure of the hirulog 3-thrombin complex and nature of the S' subsites of substrates and inhibitors.
Qiu, X., Padmanabhan, K.P., Carperos, V.E., Tulinsky, A., Kline, T., Maraganore, J.M., Fenton 2nd., J.W.(1992) Biochemistry 31: 11689-11697
- PubMed: 1445905 Search on PubMed
- DOI: https://doi.org/10.1021/bi00162a004
- Primary Citation Related Structures:
1ABI, 1ABJ - PubMed Abstract:
The X-ray crystallographic structure of the human alpha-thrombin complex with hirulog 3 (a potent, noncleavable hirudin-based peptide of the "hirulog" class containing a beta-homoarginine at the scissile bond), which is isomorphous with that of the hirugen-thrombin crystal structure, was solved at 2.3-A resolution by starting with a model for thrombin derived from the hirugen-thrombin complex and was refined by restrained least squares methods (R = 0.132). Residues of hirulog 3 were well-defined in the electron density, which included most of the pentaglycine linker and the C-terminal helical turn that was disordered in a related structure of thrombin with hirulog 1. The interactions of D-Phe1'-Pro2'-beta-homoArg3' with the active site of thrombin were essentially identical to those of related structures of PPACK- (D-Phe-Pro-Arg chloromethyl ketone) and hirulog 1-thrombin, with the guanidinium function of the arginyl P1 residue forming a hydrogen-bonding ion pair with Asp189 of the S1 site. A noticeable shift in the CA atom of beta-homoArg3' due to the methylene insertion displaces the scissile bond from attack by Ser195, thus imparting proteolytic stability to the beta-homoArg hirulog derivative. Resolution of the pentaglycine spacer, linking N- and C-terminal functional domains into a single oligopeptide bivalent inhibitor, permitted delineation of corresponding S' subsites of thrombin. The position of Gly4' (P1') is stabilized by three hydrogen bonds with His57, Lys60F, and Ser195, while the conformational angles maintained in a strained, nonallowed configuration for non-glycyl amino acids.(ABSTRACT TRUNCATED AT 250 WORDS)
- Department of Chemistry, Michigan State University, East Lansing 48824-1322.
Organizational Affiliation: