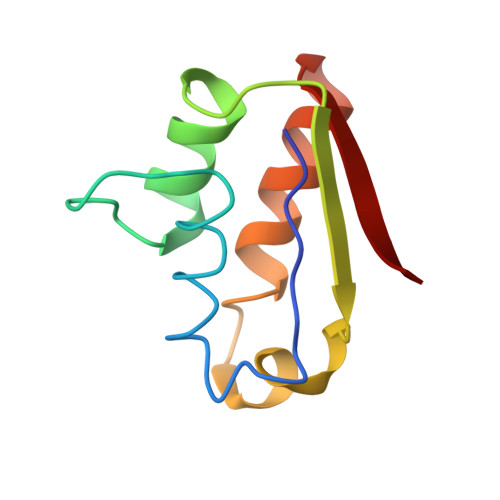
NMR 15N relaxation and structural studies reveal slow conformational exchange in barstar C40/82A.
Wong, K.B., Fersht, A.R., Freund, S.M.(1997) J Mol Biology 268: 494-511
- PubMed: 9159486 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.0989
- Primary Citation Related Structures:
1AB7 - PubMed Abstract:
Barstar an 89-residue protein consisting of four helices and a three-stranded parallel beta-sheet, is the intracellular inhibitor of the endoribonuclease barnase. Barstar C40/82A, a mutant in which the two cysteine residues have been replaced by alanine, has been used as a pseudo wild-type in folding studies and in the crystal structure of the barnase:barstar C40/82A complex. We have determined a high resolution solution structure of barstar C40/82A. The structures of barstar C40/82A and the wild-type are superimposable. A comparison with the crystal structure of the barnase:barstar C40/82A complex revealed subtle differences in the regions involved in the binding of barstar to barnase. Side-chain rotations of residues Asn33, Asp35 and Asp39 and a movement of the binding loop (Pro27-Glu32) towards the binding site of barnase facilitate the formation of interface hydrogen bonds and aromatic contacts in the complex. Extreme line broadening and missing signals in 1H-15N correlation spectra indicate substantial conformational exchange for a large subset of residues. 15N relaxation data at two magnetic field strengths, 11.74 T and 14.10 T, were used to estimate exchange contributions and to map the spectral density function at five frequencies: 0, 50, 60, 450 and 540 MHz. Based on these results, model-free calculations with the inclusion of estimated exchange contributions were used to derive order parameters and internal correlation times. The validity of this approach has been investigated with model-free calculations that incorporate longitudinal relaxation rates and heteronuclear 1H-15N NOE data only at 11.74 T and 14.10 T. The relaxation data suggest substantial conformational exchange in regions of barstar C40/82A, including the binding loop, the second and the third helices, and the second and the third strands. Amide proton exchange experiments suggest a stable hydrogen bond network for all helices and sheets except the third helix and the C-terminal of the second and the third strands. The combined results indicate a rigid body movement of the second helix and twisting motions of the beta-sheet of barstar, which might be important for the interaction with barnase.
- MRC Unit for Protein Function and Design, Cambridge Centre for Protein Engineering, University Chemical Laboratory, UK.
Organizational Affiliation: