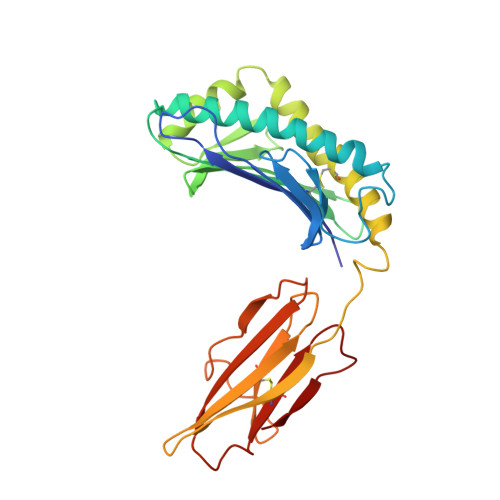
Crystal structure of the hemochromatosis protein HFE and characterization of its interaction with transferrin receptor.
Lebron, J.A., Bennett, M.J., Vaughn, D.E., Chirino, A.J., Snow, P.M., Mintier, G.A., Feder, J.N., Bjorkman, P.J.(1998) Cell 93: 111-123
- PubMed: 9546397 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81151-4
- Primary Citation Related Structures:
1A6Z - PubMed Abstract:
HFE is an MHC-related protein that is mutated in the iron-overload disease hereditary hemochromatosis. HFE binds to transferrin receptor (TfR) and reduces its affinity for iron-loaded transferrin, implicating HFE in iron metabolism. The 2.6 A crystal structure of HFE reveals the locations of hemochromatosis mutations and a patch of histidines that could be involved in pH-dependent interactions. We also demonstrate that soluble TfR and HFE bind tightly at the basic pH of the cell surface, but not at the acidic pH of intracellular vesicles. TfR:HFE stoichiometry (2:1) differs from TfR:transferrin stoichiometry (2:2), implying a different mode of binding for HFE and transferrin to TfR, consistent with our demonstration that HFE, transferrin, and TfR form a ternary complex.
- Division of Biology, California Institute of Technology, Pasadena 91125, USA.
Organizational Affiliation: