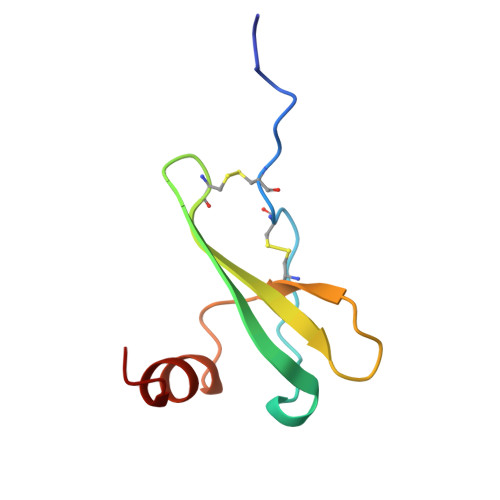
Crystal structure of chemically synthesized [N33A] stromal cell-derived factor 1alpha, a potent ligand for the HIV-1 "fusin" coreceptor.
Dealwis, C., Fernandez, E.J., Thompson, D.A., Simon, R.J., Siani, M.A., Lolis, E.(1998) Proc Natl Acad Sci U S A 95: 6941-6946
- PubMed: 9618518 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.95.12.6941
- Primary Citation Related Structures:
1A15 - PubMed Abstract:
Stromal cell-derived factor-1alpha (SDF-1alpha ) is a member of the chemokine superfamily and functions as a growth factor and chemoattractant through activation of CXCR4/LESTR/Fusin, a G protein-coupled receptor. This receptor also functions as a coreceptor for T-tropic syncytium-inducing strains of HIV-1. SDF-1alpha antagonizes infectivity of these strains by competing with gp120 for binding to the receptor. The crystal structure of a variant SDF-1alpha ([N33A]SDF-1alpha ) prepared by total chemical synthesis has been refined to 2.2-A resolution. Although SDF-1alpha adopts a typical chemokine beta-beta-beta-alpha topology, the packing of the alpha-helix against the beta-sheet is strikingly different. Comparison of SDF-1alpha with other chemokine structures confirms the hypothesis that SDF-1alpha may be either an ancestral protein from which all other chemokines evolved or the chemokine that is the least divergent from a primordial chemokine. The structure of SDF-1alpha reveals a positively charged surface ideal for binding to the negatively charged extracellular loops of the CXCR4 HIV-1 coreceptor. This ionic complementarity is likely to promote the interaction of the mobile N-terminal segment of SDF-1alpha with interhelical sites of the receptor, resulting in a biological response.
- Department of Pharmacology, Yale University School of Medicine, New Haven CT 06510, USA.
Organizational Affiliation: