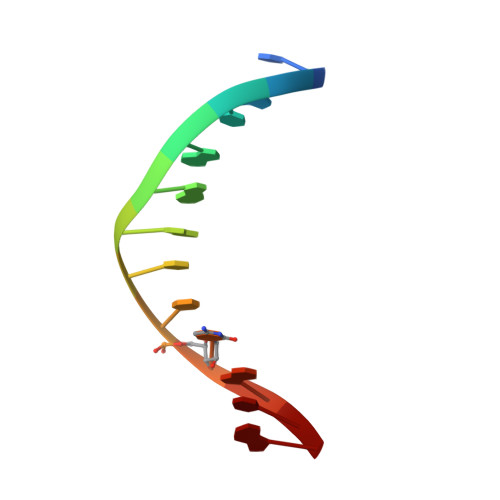
Solution structure of a DNA dodecamer containing the anti-neoplastic agent arabinosylcytosine: combined use of NMR, restrained molecular dynamics, and full relaxation matrix refinement.
Schweitzer, B.I., Mikita, T., Kellogg, G.W., Gardner, K.H., Beardsley, G.P.(1994) Biochemistry 33: 11460-11475
- PubMed: 7918360 Search on PubMed
- Primary Citation Related Structures:
170D, 171D - PubMed Abstract:
The effect of araC incorporation into the dodecamer duplex [d(CGCGAATT) (araC)d(GCG)]2 was examined by comparing its nuclear magnetic resonance (NMR)-determined solution structure with that of the control duplex d[(CGCGAATTCGCG)]2. 1H and 31P resonances in both duplexes were assigned using a combination of 2-D 1H NMR and a 3-D 31P-1H heteroTOCSY-NOESY experiment. Proton-proton distances (determined from NOESY data) and sugar dihedral angles (from NOESY and COSY data) were used in restrained molecular dynamics simulations starting from canonical A- or B-form DNA models. Both the control and araC sets of simulations converged to B-type structures. These structures were subjected to full relaxation matrix refinement to produce final structures which were in excellent agreement (R1/6 < 0.05) with the observed NOE intensities. A detailed comparison of the final control and araC structures revealed a global similarity (overall RMSD approximately 1.3 A), with significant differences localized at the araC site and neighboring bases. These included changes in sugar pucker, backbone torsion angles, base stacking, and other helical parameters. These findings are in general agreement with the previously published X-ray structure of a decamer duplex containing araC. One intriguing feature of the NMR solution structure not found in the crystal structure is the presence of an intramolecular hydrogen bond between the 2' hydroxyl on the araC sugar and the 3' phosphate group.
- Department of Pediatrics, Yale University School of Medicine, New Haven, Connecticut.
Organizational Affiliation: