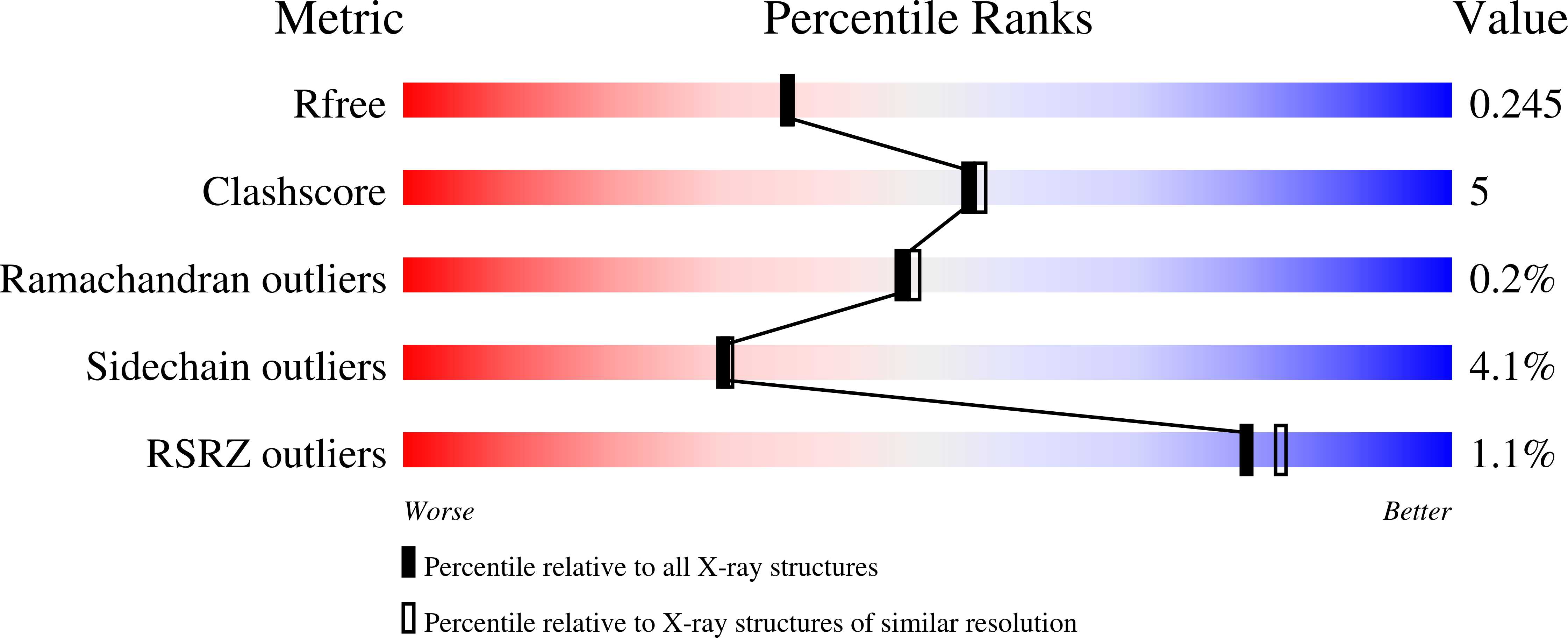
2-Aminopyridines with a shortened amino sidechain as potent, selective, and highly permeable human neuronal nitric oxide synthase inhibitors.
Vasu, D., Li, H., Hardy, C.D., Poulos, T.L., Silverman, R.B.(2022) Bioorg Med Chem 69: 116878-116878
- PubMed: 35772285
- DOI: https://doi.org/10.1016/j.bmc.2022.116878
- Primary Citation of Related Structures:
7TS1, 7TS2, 7TS3, 7TS4, 7TS5, 7TS6, 7TS7, 7TS8, 7TS9, 7TSA, 7TSB, 7TSC, 7TSD, 7TSE, 7TSF, 7TSG, 7TSH, 7TSI, 7TSK, 7TSL, 7TSM, 7TSN, 7TSO, 7TSP, 7UAM, 7UAN, 7UAO, 7US7, 7US8 - PubMed Abstract:
A series of potent, selective, and highly permeable human neuronal nitric oxide synthase inhibitors (hnNOS) based on the 2-aminopyridine scaffold with a shortened amino sidechain is reported. A rapid and simple protocol was developed to access these inhibitors in excellent yields. Neuronal nitric oxide synthase (nNOS) is a novel therapeutic target for the treatment of various neurological disorders. The major challenges in designing nNOS inhibitors in humans focus on potency, selectivity over other isoforms of nitric oxide synthases (NOSs), and blood-brain barrier permeability. In this context, we discovered a promising inhibitor, 6-(3-(4,4-difluoropiperidin-1-yl)propyl)-4-methylpyridin-2-amine dihydrochloride, that exhibits excellent potency for rat (K i = 46 nM) and human nNOS (K i = 48 nM), respectively, with 388-fold human eNOS and 135-fold human iNOS selectivity. It also displayed excellent permeability (P e = 17.3 × 10 -6 cm s -1 ) through a parallel artificial membrane permeability assay, a model for blood-brain permeability. We found that increasing lipophilicity by incorporation of fluorine atoms on the backbone of the inhibitors significantly increased potential blood-brain barrier permeability. In addition to measuring potency, isoform selectivity, and permeability of NOS inhibitors, we also explored structure-activity relationships via structures of key inhibitors complexed to various isoforms of nitric oxide synthases.
Organizational Affiliation:
Department of Chemistry, Department of Molecular Biosciences, Chemistry of Life Processes Institute, Center for Developmental Therapeutics, Northwestern University, 2145 Sheridan Road, Evanston, IL 60208-3113, United States.