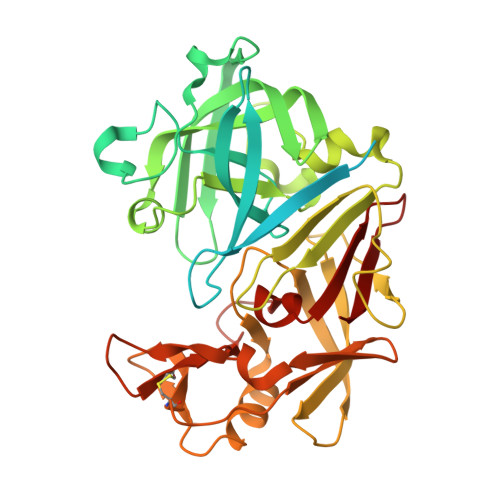
Probing ligand binding of endothiapepsin by `temperature-resolved' macromolecular crystallography.
Huang, C.Y., Aumonier, S., Engilberge, S., Eris, D., Smith, K.M.L., Leonarski, F., Wojdyla, J.A., Beale, J.H., Buntschu, D., Pauluhn, A., Sharpe, M.E., Metz, A., Olieric, V., Wang, M.(2022) Acta Crystallogr D Struct Biol 78: 964-974
- PubMed: 35916221
- DOI: https://doi.org/10.1107/S205979832200612X
- Primary Citation of Related Structures:
7QLT, 7QLU, 7QLV, 7QLW, 7QLX, 7QLY, 7QLZ, 7QM0, 7QM1, 7QM3, 7QM4, 7QM5, 7QM6, 7QM7, 7QM8, 7QM9, 7QMA, 7QMB, 7QMC, 7QMD, 7QME, 7QMF, 7QMG, 7QMH, 7QMI, 7QMJ, 7QMK, 7QML, 7QMM, 7QMN, 7QMO, 7QMP, 7QMQ, 7QMR, 7QMS, 7QMT, 7QMU, 7QMV, 7QMW, 7QMX, 7QMY, 7QMZ, 7QN0, 7QN1, 7QN2, 7QN3, 7QN4 - PubMed Abstract:
Continuous developments in cryogenic X-ray crystallography have provided most of our knowledge of 3D protein structures, which has recently been further augmented by revolutionary advances in cryoEM. However, a single structural conformation identified at cryogenic temperatures may introduce a fictitious structure as a result of cryogenic cooling artefacts, limiting the overview of inherent protein physiological dynamics, which play a critical role in the biological functions of proteins. Here, a room-temperature X-ray crystallographic method using temperature as a trigger to record movie-like structural snapshots has been developed. The method has been used to show how TL00150, a 175.15 Da fragment, undergoes binding-mode changes in endothiapepsin. A surprising fragment-binding discrepancy was observed between the cryo-cooled and physiological temperature structures, and multiple binding poses and their interplay with DMSO were captured. The observations here open up new promising prospects for structure determination and interpretation at physiological temperatures with implications for structure-based drug discovery.
Organizational Affiliation:
Swiss Light Source, Paul Scherrer Institut, Forschungsstrasse 111, Villigen-PSI, 5232 Villigen, Switzerland.