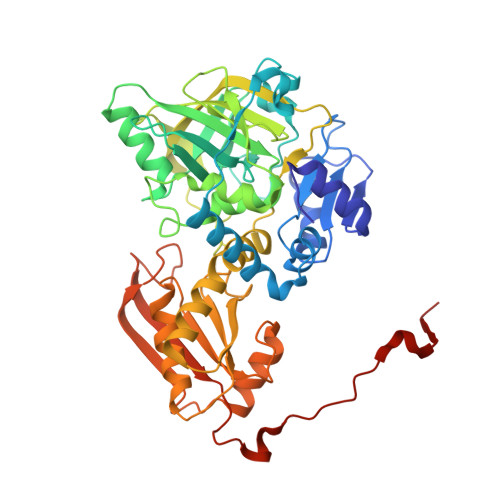
Structural and biochemical characterization of the prenylated flavin mononucleotide-dependent indole-3-carboxylic acid decarboxylase.
Gahloth, D., Fisher, K., Payne, K.A.P., Cliff, M., Levy, C., Leys, D.(2022) J Biol Chem 298: 101771-101771
- PubMed: 35218772
- DOI: https://doi.org/10.1016/j.jbc.2022.101771
- Primary Citation of Related Structures:
7P9Q - PubMed Abstract:
The ubiquitous UbiD family of reversible decarboxylases is implicated in a wide range of microbial processes and depends on the prenylated flavin mononucleotide cofactor for catalysis. However, only a handful of UbiD family members have been characterized in detail, and comparison between these has suggested considerable variability in enzyme dynamics and mechanism linked to substrate specificity. In this study, we provide structural and biochemical insights into the indole-3-carboxylic acid decarboxylase, representing an UbiD enzyme activity distinct from those previously studied. Structural insights from crystal structure determination combined with small-angle X-ray scattering measurements reveal that the enzyme likely undergoes an open-closed transition as a consequence of domain motion, an event that is likely coupled to catalysis. We also demonstrate that the indole-3-carboxylic acid decarboxylase can be coupled with carboxylic acid reductase to produce indole-3-carboxyaldehyde from indole + CO 2 under ambient conditions. These insights provide further evidence for a common mode of action in the widespread UbiD enzyme family.
Organizational Affiliation:
Manchester Institute of Biotechnology, University of Manchester, Manchester, UK.