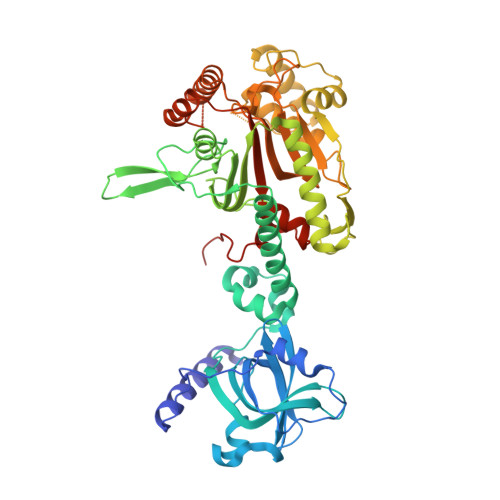
Atomic Resolution Analyses of Isocoumarin Derivatives for Inhibition of Lysyl-tRNA Synthetase.
Zhou, J., Zheng, L., Hei, Z., Li, W., Wang, J., Yu, B., Fang, P.(2020) ACS Chem Biol 15: 1016-1025
- PubMed: 32195573
- DOI: https://doi.org/10.1021/acschembio.0c00032
- Primary Citation of Related Structures:
6KA6, 6KAB, 6KBF, 6KCN, 6KCT - PubMed Abstract:
Aminoacyl-tRNA synthetases, the essential enzyme family for protein translation, are attractive targets for developing antibacterial, antifungal, and antiparasitic agents and for treating other human diseases. The antimalarial natural product cladosporin was discovered recently as a novel lysyl-tRNA synthetase (LysRS) specific inhibitor. Here, we report a thorough analysis of cladosporin derivatives using chemical synthesis, biophysical, and biochemical experiments. A series of isocoumarin derivatives with only one nonhydrogen atom/bond change per compound was synthesized. These changes include replacements of methyltetrahydropyran moiety by methylcyclohexane or cyclohexane, lactone by lactam, hydroxyl groups by methoxyl groups, and dismission of the chiral center at C3 with a Δ 3,4 double bond. We evaluated these compounds by thermal shift assays and enzymatic experiments and further studied their molecular recognition by the Plasmodium falciparum LysRS through total five high-resolution crystal structures. Our results showed that the methyltetrahydropyran moiety of cladosporin could be replaced by a more stable methylcyclohexane without reducing binding ability. Removing the methyl group from the methylcyclohexane moiety slightly decreased the interaction with LysRS. Besides, the replacement with a lactam group or a conjugated Δ 3,4 double bond within the scaffold could be two more options to optimize the compound. Lastly, the two phenolic hydroxyl groups were critical for the compounds to bind LysRS. The detailed analyses at atomic resolution in this study provide a foundation for the further development of new antibiotics from cladosporin derivatives.
Organizational Affiliation:
State Key Laboratory of Bioorganic and Natural Products Chemistry, Center for Excellence in Molecular Synthesis, Shanghai Institute of Organic Chemistry, University of Chinese Academy of Sciences, Chinese Academy of Sciences, 345 Lingling Road, Shanghai 200032, China.