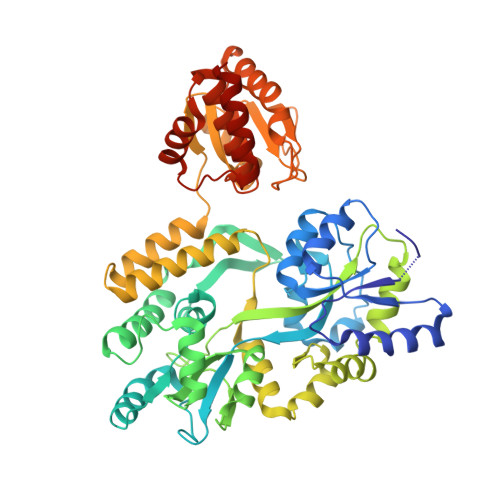
Crystal structure of the human dual specificity phosphatase 1 catalytic domain.
Gumpena, R., Lountos, G.T., Raran-Kurussi, S., Tropea, J.E., Cherry, S., Waugh, D.S.(2018) Protein Sci 27: 561-567
- PubMed: 29052270
- DOI: https://doi.org/10.1002/pro.3328
- Primary Citation of Related Structures:
6APX - PubMed Abstract:
The dual specificity phosphatase DUSP1 was the first mitogen activated protein kinase phosphatase (MKP) to be identified. It dephosphorylates conserved tyrosine and threonine residues in the activation loops of mitogen activated protein kinases ERK2, JNK1 and p38-alpha. Here, we report the crystal structure of the human DUSP1 catalytic domain at 2.49 Å resolution. Uniquely, the protein was crystallized as an MBP fusion protein in complex with a monobody that binds to MBP. Sulfate ions occupy the phosphotyrosine and putative phosphothreonine binding sites in the DUSP1 catalytic domain.
Organizational Affiliation:
Macromolecular Crystallography Laboratory, Center for Cancer Research, National Cancer Institute at Frederick, Frederick, MD, 21702.