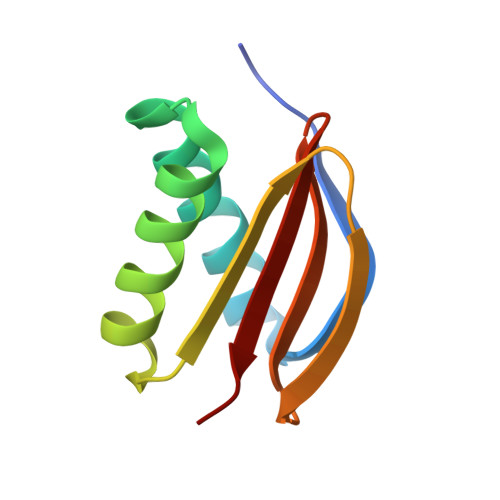
High-affinity heterotetramer formation between the large myelin-associated glycoprotein and the dynein light chain DYNLL1.
Myllykoski, M., Eichel, M.A., Jung, R.B., Kelm, S., Werner, H.B., Kursula, P.(2018) J Neurochem 147: 764-783
- PubMed: 30261098
- DOI: https://doi.org/10.1111/jnc.14598
- Primary Citation of Related Structures:
6GZJ, 6GZL - PubMed Abstract:
The close association of myelinated axons and their myelin sheaths involves numerous intercellular molecular interactions. For example, myelin-associated glycoprotein (MAG) mediates myelin-to-axon adhesion and signalling via molecules on the axonal surface. However, knowledge about intracellular binding partners of myelin proteins, including MAG, has remained limited. The two splice isoforms of MAG, S- and L-MAG, display distinct cytoplasmic domains and spatiotemporal expression profiles. We used yeast two-hybrid screening to identify interaction partners of L-MAG and found the dynein light chain DYNLL1 (also termed dynein light chain 8). DYNLL1 homodimers are known to facilitate dimerization of target proteins. L-MAG and DYNLL1 associate with high affinity, as confirmed with recombinant proteins in vitro. Structural analyses of the purified complex indicate that the DYNLL1-binding segment is localized close to the L-MAG C terminus, next to the Fyn kinase Tyr phosphorylation site. The crystal structure of the complex between DYNLL1 and its binding segment on L-MAG shows 2 : 2 binding in a parallel arrangement, indicating a heterotetrameric complex. The homology between L-MAG and previously characterized DYNLL1-ligands is limited, and some details of binding site interactions are unique for L-MAG. The structure of the complex between the entire L-MAG cytoplasmic domain and DYNLL1, as well as that of the extracellular domain of MAG, were modelled based on small-angle X-ray scattering data, allowing structural insights into L-MAG interactions on both membrane surfaces. Our data imply that DYNLL1 dimerizes L-MAG, but not S-MAG, through the formation of a specific 2 : 2 heterotetramer. This arrangement is likely to affect, in an isoform-specific manner, the functions of MAG in adhesion and myelin-to-axon signalling. OPEN SCIENCE BADGES: This article has received a badge for *Open Materials* because it provided all relevant information to reproduce the study in the manuscript. The complete Open Science Disclosure form for this article can be found at the end of the article. More information about the Open Practices badges can be found at https://cos.io/our-services/open-science-badges/. Read the Editorial Highlight for this article on page 712.
Organizational Affiliation:
Faculty of Biochemistry and Molecular Medicine, University of Oulu, Oulu, Finland.