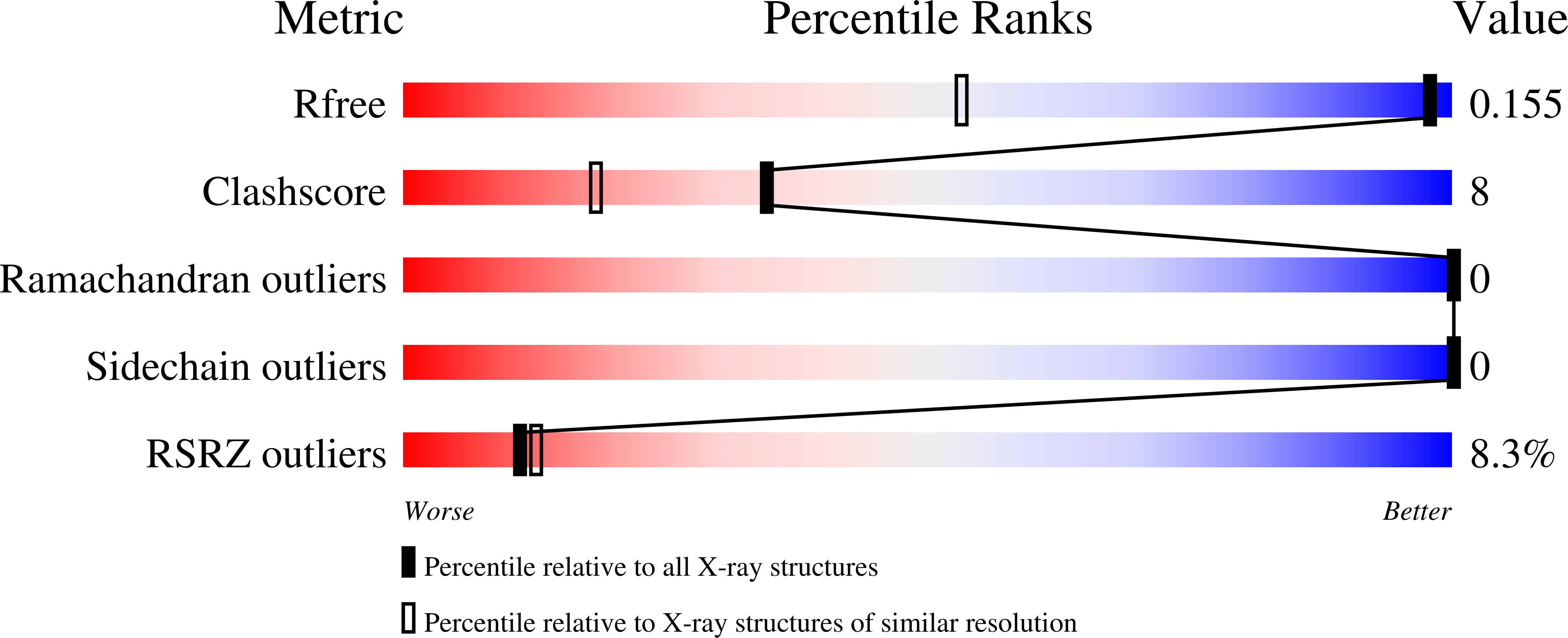
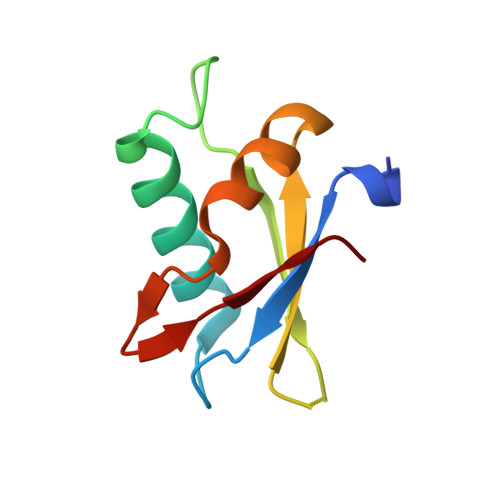
Cancer-Associated Mutations Mapped on High-Resolution Structures of the U2AF2 RNA Recognition Motifs.
Glasser, E., Agrawal, A.A., Jenkins, J.L., Kielkopf, C.L.(2017) Biochemistry 56: 4757-4761
- PubMed: 28850223
- DOI: https://doi.org/10.1021/acs.biochem.7b00551
- Primary Citation of Related Structures:
5W0G, 5W0H - PubMed Abstract:
Acquired point mutations of pre-mRNA splicing factors recur among cancers, leukemias, and related neoplasms. Several studies have established that somatic mutations of a U2AF1 subunit, which normally recognizes 3' splice site junctions, recur among myelodysplastic syndromes. The U2AF2 splicing factor recognizes polypyrimidine signals that precede most 3' splice sites as a heterodimer with U2AF1. In contrast with those of the well-studied U2AF1 subunit, descriptions of cancer-relevant U2AF2 mutations and their structural relationships are lacking. Here, we survey databases of cancer-associated mutations and identify recurring missense mutations in the U2AF2 gene. We determine ultra-high-resolution structures of the U2AF2 RNA recognition motifs (RRM1 and RRM2) at 1.1 Å resolution and map the structural locations of the mutated U2AF2 residues. Comparison with prior, lower-resolution structures of the tandem U2AF2 RRMs in the RNA-bound and apo states reveals clusters of cancer-associated mutations at the U2AF2 RRM-RNA or apo-RRM1-RRM2 interfaces. Although the role of U2AF2 mutations in malignant transformation remains uncertain, our results show that cancer-associated mutations correlate with functionally important surfaces of the U2AF2 splicing factor.
Organizational Affiliation:
Center for RNA Biology and Department of Biochemistry and Biophysics, University of Rochester School of Medicine and Dentistry , Rochester, New York 14642, United States.