Structures of human O-GlcNAcase and its complexes reveal a new substrate recognition mode.
Li, B., Li, H., Lu, L., Jiang, J.(2017) Nat Struct Mol Biol 24: 362-369
- PubMed: 28319083
- DOI: https://doi.org/10.1038/nsmb.3390
- Primary Citation of Related Structures:
5TKE, 5UN8, 5UN9 - PubMed Abstract:
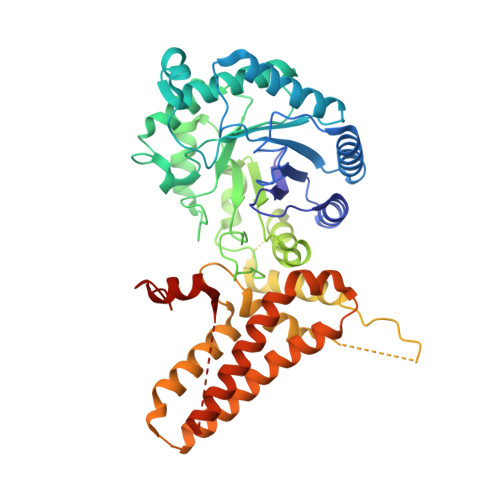
Human O-GlcNAcase (hOGA) is the unique enzyme responsible for the hydrolysis of the O-linked β-N-acetyl glucosamine (O-GlcNAc) modification, an essential protein glycosylation event that modulates the function of numerous cellular proteins in response to nutrients and stress. Here we report crystal structures of a truncated hOGA, which comprises the catalytic and stalk domains, in apo form, in complex with an inhibitor, and in complex with a glycopeptide substrate. We found that hOGA forms an unusual arm-in-arm homodimer in which the catalytic domain of one monomer is covered by the stalk domain of the sister monomer to create a substrate-binding cleft. Notably, the residues on the cleft surface afford extensive interactions with the peptide substrate in a recognition mode that is distinct from that of its bacterial homologs. These structures represent the first model of eukaryotic enzymes in the glycoside hydrolase 84 (GH84) family and provide a crucial starting point for understanding the substrate specificity of hOGA, which regulates a broad range of biological and pathological processes.
Organizational Affiliation:
Pharmaceutical Sciences Division, School of Pharmacy, University of Wisconsin-Madison, Madison, Wisconsin, USA.