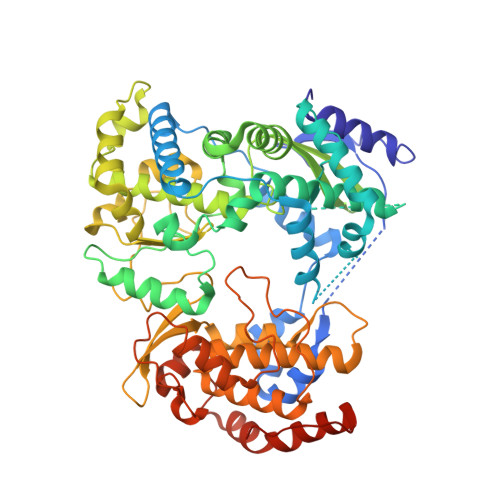
Targeting flavivirus RNA dependent RNA polymerase through a pyridobenzothiazole inhibitor.
Tarantino, D., Cannalire, R., Mastrangelo, E., Croci, R., Querat, G., Barreca, M.L., Bolognesi, M., Manfroni, G., Cecchetti, V., Milani, M.(2016) Antiviral Res 134: 226-235
- PubMed: 27649989
- DOI: https://doi.org/10.1016/j.antiviral.2016.09.007
- Primary Citation of Related Structures:
5IQ6 - PubMed Abstract:
RNA dependent RNA polymerases (RdRp) are essential enzymes for flavivirus replication. Starting from an in silico docking analysis we identified a pyridobenzothiazole compound, HeE1-2Tyr, able to inhibit West Nile and Dengue RdRps activity in vitro, which proved effective against different flaviviruses in cell culture. Crystallographic data show that HeE1-2Tyr binds between the fingers domain and the priming loop of Dengue virus RdRp (Site 1). Conversely, enzyme kinetics, binding studies and mutational analyses suggest that, during the catalytic cycle and assembly of the RdRp-RNA complex, HeE1-2Tyr might be hosted in a distinct binding site (Site 2). RdRp mutational studies, driven by in silico docking analysis, allowed us to locate the inhibition Site 2 in the thumb domain. Taken together, our results provide innovative concepts for optimization of a new class of anti-flavivirus compounds.
Organizational Affiliation:
Dipartimento di Bioscienze, Università di Milano, Via Celoria 26, I-20133, Milano, Italy; CNR-IBF, Istituto di Biofisica, Via Celoria 26, I-20133, Milano, Italy.