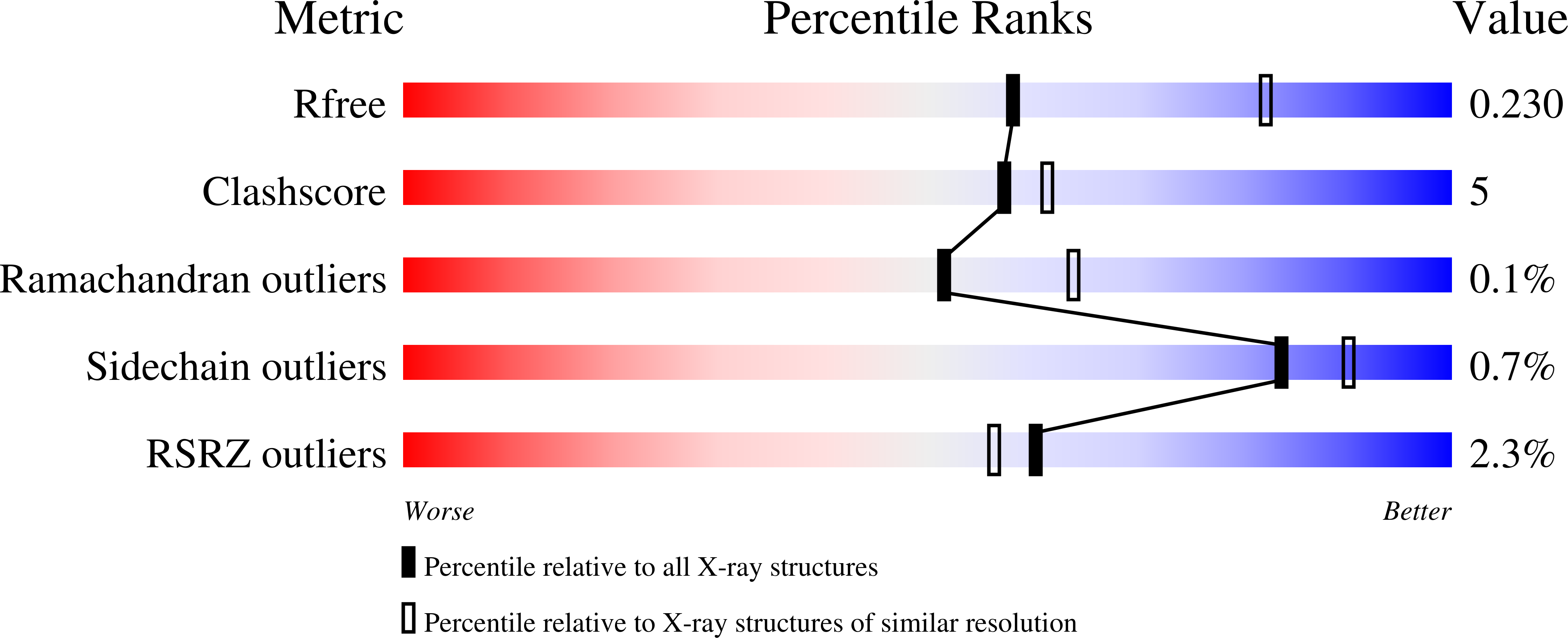
Structural Analysis of Human Kdm5B Guides Histone Demethylase Inhibitor Development.
Johansson, C., Velupillai, S., Tumber, A., Szykowska, A., Hookway, E.S., Nowak, R.P., Strain-Damerell, C., Gileadi, C., Philpott, M., Burgess-Brown, N., Wu, N., Kopec, J., Nuzzi, A., Steuber, H., Egner, U., Badock, V., Munro, S., Lathangue, N.B., Westaway, S., Brown, J., Athanasou, N., Prinjha, R., Brennan, P.E., Oppermann, U.(2016) Nat Chem Biol 12: 539
- PubMed: 27214403
- DOI: https://doi.org/10.1038/nchembio.2087
- Primary Citation of Related Structures:
4UF0, 5A1F, 5A3P, 5A3T, 5A3W, 5FPU, 5FPV, 5FUN, 5FUP, 5FV3, 5FWJ - PubMed Abstract:
Members of the KDM5 (also known as JARID1) family are 2-oxoglutarate- and Fe(2+)-dependent oxygenases that act as histone H3K4 demethylases, thereby regulating cell proliferation and stem cell self-renewal and differentiation. Here we report crystal structures of the catalytic core of the human KDM5B enzyme in complex with three inhibitor chemotypes. These scaffolds exploit several aspects of the KDM5 active site, and their selectivity profiles reflect their hybrid features with respect to the KDM4 and KDM6 families. Whereas GSK-J1, a previously identified KDM6 inhibitor, showed about sevenfold less inhibitory activity toward KDM5B than toward KDM6 proteins, KDM5-C49 displayed 25-100-fold selectivity between KDM5B and KDM6B. The cell-permeable derivative KDM5-C70 had an antiproliferative effect in myeloma cells, leading to genome-wide elevation of H3K4me3 levels. The selective inhibitor GSK467 exploited unique binding modes, but it lacked cellular potency in the myeloma system. Taken together, these structural leads deliver multiple starting points for further rational and selective inhibitor design.
Organizational Affiliation:
Structural Genomics Consortium, University of Oxford, Headington, UK.