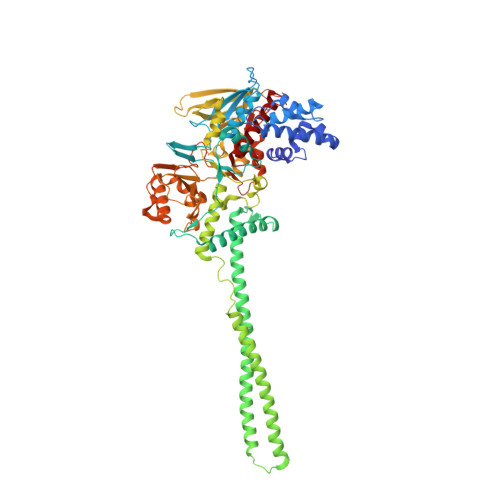
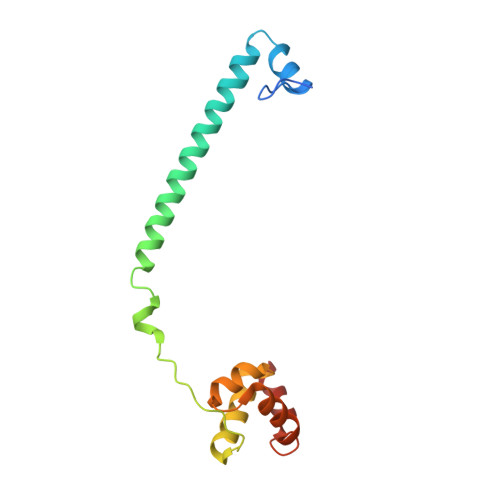
Crystal Structure of LSD1 in Complex with 4-[5-(Piperidin-4-ylmethoxy)-2-(p-tolyl)pyridin-3-yl]benzonitrile.
Niwa, H., Sato, S., Hashimoto, T., Matsuno, K., Umehara, T.(2018) Molecules 23
- PubMed: 29949906
- DOI: https://doi.org/10.3390/molecules23071538
- Primary Citation of Related Structures:
5YJB - PubMed Abstract:
Because lysine-specific demethylase 1 (LSD1) regulates the maintenance of cancer stem cell properties, small-molecule inhibitors of LSD1 are expected to be useful for the treatment of several cancers. Reversible inhibitors of LSD1 with submicromolar inhibitory potency have recently been reported, but their exact binding modes are poorly understood. In this study, we synthesized a recently reported reversible inhibitor, 4-[5-(piperidin-4-ylmethoxy)-2-( p -tolyl)pyridin-3-yl]benzonitrile, which bears a 4-piperidinylmethoxy group, a 4-methylphenyl group, and a 4-cyanophenyl group on a pyridine ring, and determined the crystal structure of LSD1 in complex with this inhibitor at 2.96 Å. We observed strong electron density for the compound, showing that its cyano group forms a hydrogen bond with Lys661, which is a critical residue in the lysine demethylation reaction located deep in the catalytic center of LSD1. The piperidine ring interacts with the side chains of Asp555 and Asn540 in two conformations, and the 4-methylphenyl group is bound in a hydrophobic pocket in the catalytic center. Our elucidation of the binding mode of this compound can be expected to facilitate the rational design of more-potent reversible LSD1 inhibitors.
Organizational Affiliation:
Laboratory for Epigenetics Drug Discovery, RIKEN Center for Biosystems Dynamics Research (BDR), 1-7-22 Suehiro-cho, Tsurumi, Yokohama 230-0045, Japan. hideaki.niwa@riken.jp.