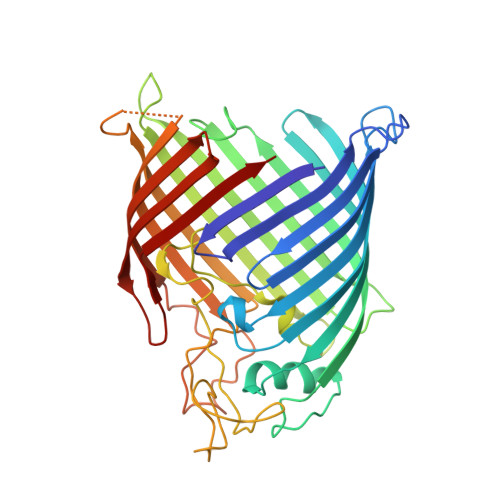
Structural and functional analysis of the beta-barrel domain of BamA from Escherichia coli.
Ni, D., Wang, Y., Yang, X., Zhou, H., Hou, X., Cao, B., Lu, Z., Zhao, X., Yang, K., Huang, Y.(2014) FASEB J 28: 2677-2685
- PubMed: 24619089
- DOI: https://doi.org/10.1096/fj.13-248450
- Primary Citation of Related Structures:
4N75 - PubMed Abstract:
In gram-negative bacteria, the assembly of outer membrane proteins (OMPs) requires a β-barrel assembly machinery (BAM) complex, of which BamA is an essential and evolutionarily conserved component. To elucidate the mechanism of BamA-mediated OMP biogenesis, we determined the crystal structure of the C-terminal transmembrane domain of BamA from Escherichia coli (EcBamA) at 2.6 Å resolution. The structure reveals 2 distinct features. First, a portion of the extracellular side of the β barrel is composed of 5 markedly short β strands, and the loops stemming from these β strands form a potential surface cavity, filled by a portion of the L6 loop that includes the conserved VRGF/Y motif found in the Omp85 family. Second, the 4 extracellular loops L3, L4, L6, and L7 of EcBamA form a dome over the barrel, stabilized by a salt-bridge interaction network. Functional data show that hydrophilic-to-hydrophobic mutations of the potential hydrophilic surface cavity and a single Arg547Ala point mutation that may destabilize the dome severely affect the function of EcBamA. Our structure of the EcBamA β barrel and structure-based mutagenesis studies suggest that the transmembrane β strands of OMP substrates may integrate into the outer membrane at the interface of the first and last β strands of the EcBamA barrel, whereas the soluble loops or domains may be transported out of the cell via the hydrophilic surface cavity on dislocation of the VRGF/Y motif of L6. In addition, the dome over the barrel may play an important role in maintaining the efficiency of OMP biogenesis.
Organizational Affiliation:
National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing, China; School of Public Health, Tianjin Medical University, Tianjin, China;