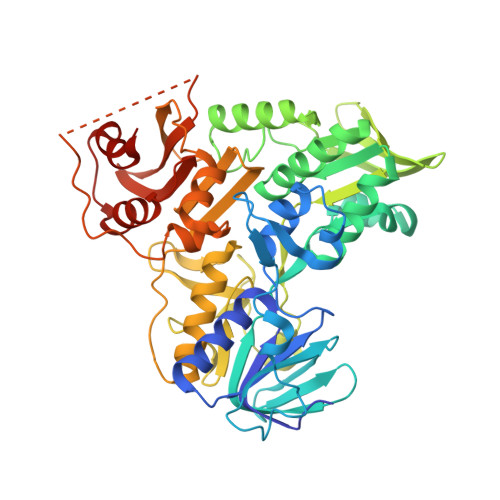
Structural Insights Into the Coenzyme Mediated Monomer-Dimer Transition of the Pro-Apoptotic Apoptosis Inducing Factor.
Ferreira, P., Villanueva, R., Martinez-Julvez, M., Herguedas, B., Marcuello, C., Fernandez-Silva, P., Cabon, L., Hermoso, J.A., Lostao, A., Susin, S.A., Medina, M.(2014) Biochemistry 53: 4204
- PubMed: 24914854
- DOI: https://doi.org/10.1021/bi500343r
- Primary Citation of Related Structures:
4BUR, 4BV6 - PubMed Abstract:
The apoptosis-inducing factor (AIF) is a mitochondrial-flavoprotein that, after cell death induction, is distributed to the nucleus to mediate chromatinolysis. In mitochondria, AIF is present in a monomer-dimer equilibrium that after reduction by NADH gets displaced toward the dimer. The crystal structure of the human AIF (hAIF):NAD(H)-bound dimer revealed one FAD and, unexpectedly, two NAD(H) molecules per protomer. A 1:2 hAIF:NAD(H) binding stoichiometry was additionally confirmed in solution by using surface plasmon resonance. The here newly discovered NAD(H)-binding site includes residues mutated in human disorders, and accommodation of the coenzyme in it requires restructuring of a hAIF portion within the 509-560 apoptogenic segment. Disruption of interactions at the dimerization surface by production of the hAIF E413A/R422A/R430A mutant resulted in a nondimerizable variant considerably less efficiently stabilizing charge-transfer complexes upon coenzyme reduction than WT hAIF. These data reveal that the coenzyme-mediated monomer-dimer transition of hAIF modulates the conformation of its C-terminal proapoptotic domain, as well as its mechanism as reductase. These observations suggest that both the mitochondrial and apoptotic functions of hAIF are interconnected and coenzyme controlled: a key information in the understanding of the physiological role of AIF in the cellular life and death cycle.
Organizational Affiliation:
Departamento de Bioquímica y Biología Molecular y Celular, ‡Instituto de Biocomputación y Física de Sistemas Complejos (BIFI)-Joint Unit BIFI-IQFR (CSIC), and §Laboratorio de Microscopias Avanzadas, Instituto de Nanociencia de Aragón (INA), Universidad de Zaragoza , Zaragoza, Spain.