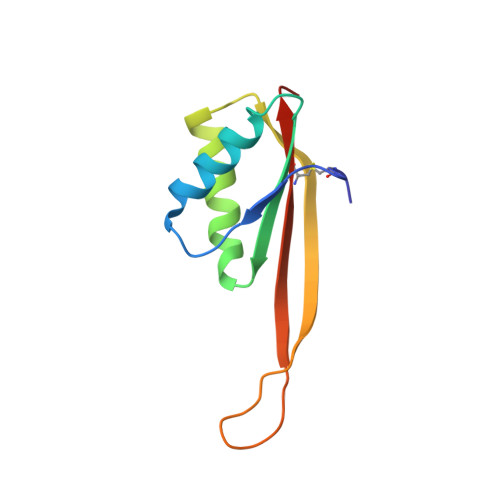
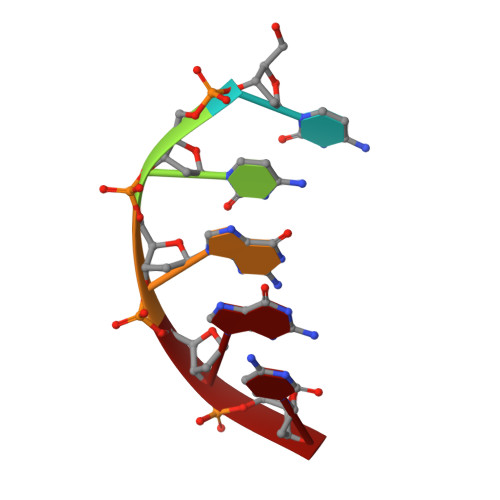
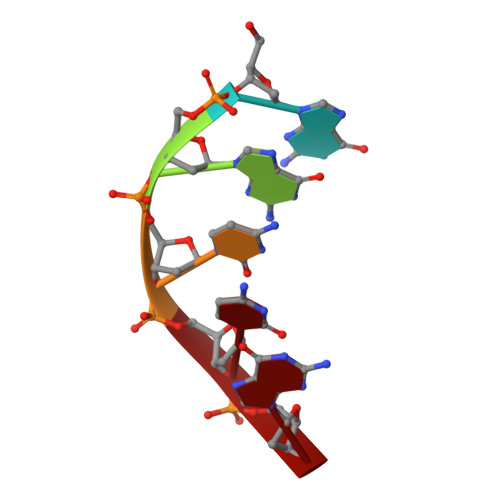
Crystal structure of archaeal chromatin protein Alba2-double-stranded DNA complex from Aeropyrum pernix K1.
Tanaka, T., Padavattan, S., Kumarevel, T.(2012) J Biol Chem 287: 10394-10402
- PubMed: 22334696
- DOI: https://doi.org/10.1074/jbc.M112.343210
- Primary Citation of Related Structures:
3U6Y - PubMed Abstract:
All thermophilic and hyperthermophilic archaea encode homologs of dimeric Alba (Sac10b) proteins that bind cooperatively at high density to DNA. Here, we report the 2.0 Å resolution crystal structure of an Alba2 (Ape10b2)-dsDNA complex from Aeropyrum pernix K1. A rectangular tube-like structure encompassing duplex DNA reveals the positively charged residues in the monomer-monomer interface of each dimer packing on either side of the bound dsDNA in successive minor grooves. The extended hairpin loop connecting strands β3 and β4 undergoes significant conformational changes upon DNA binding to accommodate the other Alba2 dimer during oligomerization. Mutational analysis of key interacting residues confirmed the specificity of Alba2-dsDNA interactions.
Organizational Affiliation:
RIKEN SPring-8 Center, Harima Institute, 1-1-1 Kouto, Sayo, Hyogo 679-5148, Japan.