Circular permutation of Bacillus circulans xylanase: a kinetic and structural study.
Reitinger, S., Yu, Y., Wicki, J., Ludwiczek, M., D'Angelo, I., Baturin, S., Okon, M., Strynadka, N.C., Lutz, S., Withers, S.G., McIntosh, L.P.(2010) Biochemistry 49: 2464-2474
- PubMed: 20163191
- DOI: https://doi.org/10.1021/bi100036f
- Primary Citation of Related Structures:
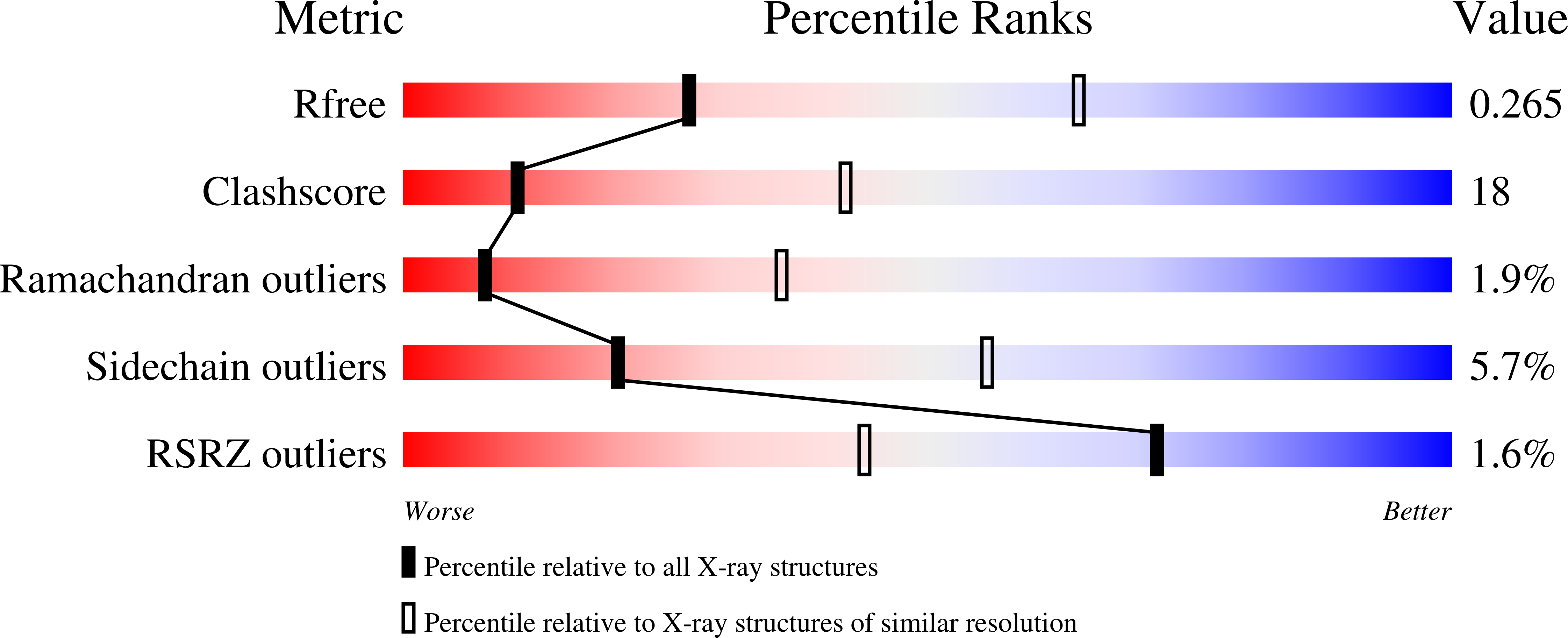
3LB9 - PubMed Abstract:
The 20 kDa Bacillus circulans Bcx is a well-studied endoxylanase with a beta-jellyroll fold that places its N- and C-termini in salt bridge contact. Initial experiments verified that Bcx could be circularly permuted by PCR methods to introduce new termini in loop regions while linking its native termini directly or via one or two glycines. Subsequently, a library of circular permutants, generated by random DNase cleavage of the circularized Bcx gene, was screened for xylanase activity on xylan in Congo Red-stained agar. Analysis of 35 unique active circular permutants revealed that, while many of the new termini were introduced in external loops as anticipated, a surprising number were also located within beta-strands. Furthermore, several permutations placed key catalytic residues at or near the new termini with minimal deleterious effects on activity and, in one case, a 4-fold increase. The structure of one permutant was determined by X-ray crystallography, whereas three others were probed by NMR spectroscopy. These studies revealed that the overall conformation of Bcx changed very little in response to circular permutation, with effects largely being limited to increased local mobility near the new and the linked old termini and to a decrease in global stability against thermal denaturation. This library of circularly permuted xylanases provides an excellent set of new start points for directed evolution of this commercially important enzyme, as well as valuable constructs for intein-mediated replacement of key catalytic residues with unnatural analogues. Such approaches should permit new insights into the mechanism of enzymatic glycoside hydrolysis.
Organizational Affiliation:
Department of Chemistry, University of British Columbia, Vancouver, British Columbia, Canada V6T 1Z1.