Identification and Analysis of Residues Contained on Beta --> Alpha Loops of the Dual-Substrate (Betaalpha)(8) Phosphoribosyl Isomerase a (Pria) Specific for its Phosphoribosyl Anthranilate Isomerase Activity.
Noda-Garcia, L., Camacho-Zarco, A.R., Verdel-Aranda, K., Wright, H., Soberon, X., Fulop, V., Barona-Gomez, F.(2010) Protein Sci 19: 535
- PubMed: 20066665
- DOI: https://doi.org/10.1002/pro.331
- Primary Citation of Related Structures:
2X30 - PubMed Abstract:
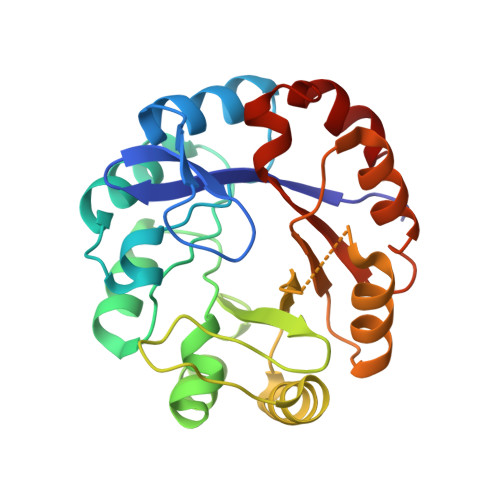
A good model to experimentally explore evolutionary hypothesis related to enzyme function is the ancient-like dual-substrate (beta alpha)(8) phosphoribosyl isomerase A (PriA), which takes part in both histidine and tryptophan biosynthesis in Streptomyces coelicolor and related organisms. In this study, we determined the Michaelis-Menten enzyme kinetics for both isomerase activities in wild-type PriA from S. coelicolor and in selected single-residue monofunctional mutants, identified after Escherichia coli in vivo complementation experiments. Structural and functional analyses of a hitherto unnoticed residue contained on the functionally important beta --> alpha loop 5, namely, Arg(139), which was postulated on structural grounds to be important for the dual-substrate specificity of PriA, is presented for the first time. Indeed, enzyme kinetics analyses done on the mutant variants PriA_Ser(81)Thr and PriA_Arg(139)Asn showed that these residues, which are contained on beta --> alpha loops and in close proximity to the N-terminal phosphate-binding site, are essential solely for the phosphoribosyl anthranilate isomerase activity of PriA. Moreover, analysis of the X-ray crystallographic structure of PriA_Arg(139)Asn elucidated at 1.95 A herein strongly implicates the occurrence of conformational changes in this beta --> alpha loop as a major structural feature related to the evolution of the dual-substrate specificity of PriA. It is suggested that PriA has evolved by tuning a fine energetic balance that allows the sufficient degree of structural flexibility needed for accommodating two topologically dissimilar substrates--within a bifunctional and thus highly constrained active site--without compromising its structural stability.
Organizational Affiliation:
Evolution of Metabolic Diversity Laboratory, Laboratorio Nacional de Genómica para la Biodiversidad (Langebio), CINVESTAV-IPN, Km 9.6 Libramiento Norte, Carretera Irapuato-León, Irapuato, C.P. 36822, México.