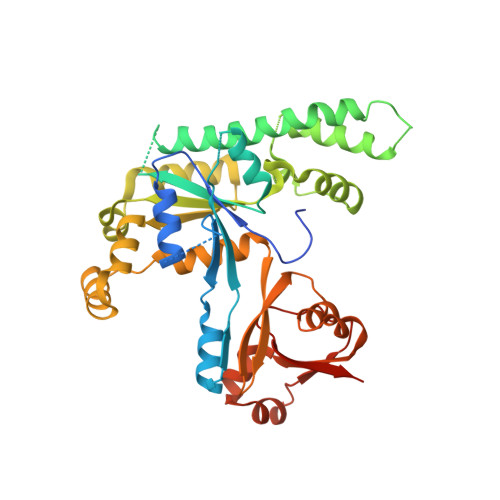
Human OLA1 defines an ATPase subfamily in the Obg family of GTP-binding proteins
Koller-Eichhorn, R., Marquardt, T., Gail, R., Wittinghofer, A., Kostrewa, D., Kutay, U., Kambach, C.(2007) J Biological Chem 282: 19928-19937
- PubMed: 17430889
- DOI: https://doi.org/10.1074/jbc.M700541200
- Primary Citation of Related Structures:
2OHF - PubMed Abstract:
Purine nucleotide-binding proteins build the large family of P-loop GTPases and related ATPases, which perform essential functions in all kingdoms of life. The Obg family comprises a group of ancient GTPases belonging to the TRAFAC (for translation factors) class and can be subdivided into several distinct protein subfamilies. The founding member of one of these subfamilies is the bacterial P-loop NTPase YchF, which had so far been assumed to act as GTPase. We have biochemically characterized the human homologue of YchF and found that it binds and hydrolyzes ATP more efficiently than GTP. For this reason, we have termed the protein hOLA1, for human Obg-like ATPase 1. Further biochemical characterization of YchF proteins from different species revealed that ATPase activity is a general but previously missed feature of the YchF subfamily of Obg-like GTPases. To explain ATP specificity of hOLA1, we have solved the x-ray structure of hOLA1 bound to the nonhydrolyzable ATP analogue AMPPCP. Our structural data help to explain the altered nucleotide specificity of YchF homologues and identify the Ola1/YchF subfamily of the Obg-related NTPases as an exceptional example of a single protein subfamily, which has evolved altered nucleotide specificity within a distinct protein family of GTPases.
- Institute of Biochemistry, ETH Zurich, 8093 Zurich, Switzerland.
Organizational Affiliation: